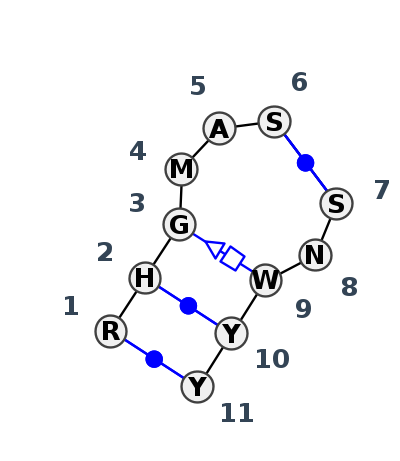
Motif IL_47352.3 Version IL_47352.3 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | break | 7 | 8 | 9 | 10 | 11 | 1-11 | 2-10 | 3-9 | 4-8 | 5-8 | 6-7 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_8B0X_062 | 8B0X | 0.4033 | 0 | Triple sheared related (H) | A | SSU rRNA | A | 1430 | A | 1431 | G | 1432 | A | 1433 | A | 1434 | G | 1435 | * | C | 1466 | C | 1467 | A | 1468 | C | 1469 | U | 1470 | cWW | ntSH | tSH | ntHS | cWW | |
| 2 | IL_9H3G_029 | 9H3G | 0.4542 | 0 | A | LSU rRNA | G | 749 | U | 750 | G | 751 | C | 752 | A | 753 | C | 754 | * | G | 736 | G | 737 | A | 738 | U | 739 | C | 740 | cWW | cwW | tSH | tHS | cWW | ||
| 3 | IL_8GLP_051 | 8GLP | 0.3741 | 0 | L5 | LSU rRNA | G | 1455 | C | 1456 | G | 1457 | C | 1458 | A | 1459 | C | 1460 | * | G | 1425 | G | 1426 | A | 1427 | U | 1428 | C | 1429 | cWW | cWW | tHS | cWW | |||
| 4 | IL_1M5K_001 | 1M5K | 0.1523 | 1 | B+A | Hairpin ribozyme | G | 6 | A | 7 | G | 8 | A | 9 | A | 10 | G | 11 | * | C | 11 | A2M | 12 | U | 14 | C | 15 | C | 16 | cWW | cWWa | tSH | ntHS | cWW | ||
| 5 | IL_1X9K_001 | 1X9K | 0.1769 | 1 | B+A | RNA (13-mer) | G | 6 | A | 7 | G | 8 | A | 9 | A | 10 | G | 11 | * | C | 5 | A | 6 | U | 8 | C | 9 | C | 10 | cWW | tSH | ntHS | cWW | |||
| 6 | IL_1ZFT_001 | 1ZFT | 0.1076 | -1 | B+A | RNA (12-mer) | G | 6 | A | 7 | I | 8 | A | 9 | A | 10 | G | 11 | * | C | 4 | A2M | 5 | U | 7 | C | 8 | C | 9 | cWW | cWWa | tSH | tHS | cWW | ||
| 7 | IL_3BBM_001 | 3BBM | 0.0000 | 0 | B+A | Loop A and Loop B Ribozyme strand | G | 6 | A | 7 | G | 8 | A | 9 | A | 10 | G | 11 | * | C | 4 | A2M | 5 | U | 7 | C | 8 | C | 9 | cWW | cWWa | tSH | ntHS | cWW | ||
| 8 | IL_2NPY_001 | 2NPY | 0.1223 | 0 | B+A | RNA (12-mer) | G | 6 | A | 7 | G | 8 | A | 9 | A | 10 | G | 11 | * | C | 4 | 2AU | 5 | U | 7 | C | 8 | C | 9 | cWW | cWWa | tSH | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GAGAAG*CGUCC | 2 |
| AAGAAG*CCACU | 1 |
| GUGCAC*GGAUC | 1 |
| GCGCAC*GGAUC | 1 |
| GAGAAG*C(A2M)GUCC | 1 |
| GAGAAG*CAGUCC | 1 |
| GAAAG*CGUCC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AGAA*GUC | 2 |
| AGAA*CAC | 1 |
| UGCA*GAU | 1 |
| CGCA*GAU | 1 |
| AGAA*(A2M)GUC | 1 |
| AGAA*AGUC | 1 |
| AAA*GUC | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 2 parents |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (7)
- Triple sheared related (H) (1)
- Basepair signature
- cWW-cWW-tSH-L-R-L-cWW
- Heat map statistics
- Min 0.11 | Avg 0.28 | Max 0.50
Coloring options: