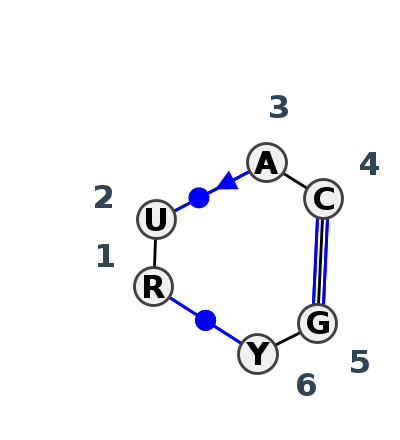
Motif IL_52092.2 Version IL_52092.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | break | 5 | 6 | 1-6 | 2-3 | 4-5 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_9H3G_109 | 9H3G | 0.1800 | 5 | A | LSU rRNA | G | 2518 | U | 2519 | A | 2521 | C | 2524 | * | G | 2583 | C | 2586 | cWW | cWS | cWW |
| 2 | IL_8GLP_135 | 8GLP | 0.1212 | 5 | L5 | LSU rRNA | G | 4082 | U | 4083 | A | 4085 | C | 4088 | * | G | 4161 | C | 4164 | cWW | cWS | cWW |
| 3 | IL_9AXU_090 | 9AXU | 0.1645 | 5 | 2 | LSU rRNA | A | 2616 | U | 2617 | A | 2619 | C | 2622 | * | G | 2679 | U | 2682 | cWW | cWS | cWW |
| 4 | IL_8OI5_098 | 8OI5 | 0.0000 | 5 | 1 | LSU rRNA | A | 2498 | U | 2499 | A | 2501 | C | 2504 | * | G | 2556 | U | 2559 | cWW | cWS | cWW |
| 5 | IL_8P9A_324 | 8P9A | 0.0882 | 5 | AR | LSU rRNA | A | 2520 | U | 2521 | A | 2523 | C | 2526 | * | G | 2584 | U | 2587 | cWW | cWS | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AUGAAGC*GGGU | 2 |
| GUGAAUC*GGGC | 1 |
| GUGAGGC*GCUC | 1 |
| AUGAAGC*GCGU | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UGAAG*GG | 2 |
| UGAAU*GG | 1 |
| UGAGG*CU | 1 |
| UGAAG*CG | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (5)
- Basepair signature
- cWW-cWS-cWW
- Heat map statistics
- Min 0.09 | Avg 0.16 | Max 0.39
Coloring options: