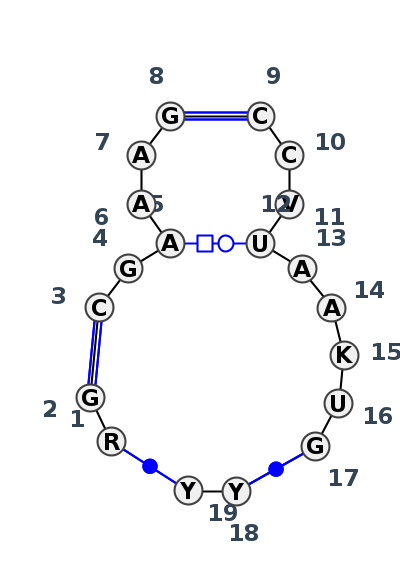
Motif J4_14595.2 Version J4_14595.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | break | 3 | 4 | 5 | 6 | 7 | 8 | break | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | break | 18 | 19 | 1-19 | 2-3 | 5-12 | 7-10 | 8-9 | 11-12 | 17-18 | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | J4_8GLP_016 | 8GLP | 0.1259 | 4 | S2 | SSU rRNA | A | 671 | G | 673 | * | C | 1032 | G | 1033 | A | 1034 | A | 1035 | A | 1036 | G | 1037 | * | C | 1078 | C | 1079 | A | 1080 | PSU | 1081 | A | 1082 | A | 1084 | G | 1086 | U | 1088 | G | 1089 | * | U | 1160 | U | 1161 | cWW | cWW | tHW | ncWW | cWW | cWW | |
| 2 | J4_8P9A_022 | 8P9A | 0.0884 | 4 | sR | SSU rRNA | A | 622 | G | 624 | * | C | 975 | G | 976 | A | 977 | A | 978 | A | 979 | G | 980 | * | C | 1021 | C | 1022 | A | 1023 | U | 1024 | A | 1025 | A | 1027 | U | 1029 | U | 1031 | G | 1032 | * | U | 1103 | U | 1104 | cWW | cWW | tHW | ncWW | cWW | cWW | |
| 3 | J4_8CRE_022 | 8CRE | 0.0791 | 4 | CM | SSU rRNA | A | 620 | G | 622 | * | C | 960 | G | 961 | A | 962 | A | 963 | A | 964 | G | 965 | * | C | 1006 | C | 1007 | A | 1008 | U | 1009 | A | 1010 | A | 1012 | U | 1014 | U | 1016 | G | 1017 | * | U | 1088 | U | 1089 | cWW | cWW | tHW | ncWW | cWW | cWW | |
| 4 | J4_9H3G_014 | 9H3G | 0.0000 | 4 | h1 | SSU rRNA | A | 624 | G | 626 | * | C | 976 | G | 977 | A | 978 | A | 979 | A | 980 | G | 981 | * | C | 1022 | C | 1023 | A | 1024 | PSU | 1025 | A | 1026 | A | 1028 | G | 1030 | U | 1032 | G | 1033 | * | PSU | 1104 | U | 1105 | cWW | cWW | tHW | ncWW | cWW | ncSH | cWW |
| 5 | J4_8B0X_003 | 8B0X | 0.0670 | 4 | A | SSU rRNA | G | 575 | G | 577 | * | C | 764 | G | 765 | A | 766 | A | 767 | A | 768 | G | 769 | * | C | 810 | C | 811 | G | 812 | U | 813 | A | 814 | A | 816 | G | 818 | U | 820 | G | 821 | * | C | 879 | C | 880 | cWW | cWW | tHW | ncWW | cWW | ncSH | cWW |
| 6 | J4_4LFB_003 | 4LFB | 0.0926 | 4 | A | SSU rRNA | G | 575 | G | 577 | * | C | 764 | G | 765 | A | 766 | A | 767 | A | 768 | G | 769 | * | C | 810 | C | 811 | C | 812 | U | 813 | A | 814 | A | 816 | G | 818 | U | 820 | G | 821 | * | C | 879 | C | 880 | cWW | cWW | tHW | ncWW | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AAG*CGAAAG*CCAUAAACUAUG*UU | 2 |
| AAG*CGAAAG*CCA(PSU)AAACGAUG*UU | 1 |
| AAG*CGAAAG*CCA(PSU)AAACGAUG*PSUU | 1 |
| GCG*CGAAAG*CCGUAAACGAUG*CC | 1 |
| GGG*CGAAAG*CCCUAAACGAUG*CC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| A*GAAA*CA(PSU)AAACGAU* | 2 |
| A*GAAA*CAUAAACUAU* | 2 |
| C*GAAA*CGUAAACGAU* | 1 |
| G*GAAA*CCUAAACGAU* | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (6)
- Basepair signature
- cWW-cWW-cWW-F-F-tHW-F-F-F-F-F-cWW-F-F
- Heat map statistics
- Min 0.05 | Avg 0.09 | Max 0.16
Coloring options: