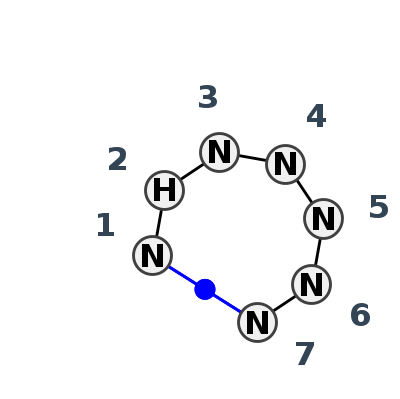
Motif HL_31585.4 Version HL_31585.4 of this group appears in releases 3.98 to 3.99
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 1-7 | 2-5 | 2-6 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_4V9F_063 | 4V9F | 0.7245 | 1 | GNRA variation | 0 | LSU rRNA | U | 2563 | C | 2565 | A | 2566 | G | 2567 | A | 2568 | A | 2569 | G | 2570 | cWW | ||
| 2 | HL_7A0S_060 | 7A0S | 0.7258 | 1 | GNRA variation | X | LSU rRNA | U | 2507 | A | 2509 | A | 2510 | G | 2511 | A | 2512 | A | 2513 | G | 2514 | cWW | ||
| 3 | HL_4WF9_057 | 4WF9 | 0.7725 | 1 | GNRA variation | X | LSU rRNA | U | 2555 | U | 2557 | A | 2558 | G | 2559 | U | 2560 | C | 2561 | G | 2562 | cWW | ||
| 4 | HL_5J7L_194 | 5J7L | 0.7462 | 1 | GNRA variation | DA | LSU rRNA | U | 2528 | A | 2530 | A | 2531 | G | 2532 | U | 2533 | A | 2534 | G | 2535 | cWW | ||
| 5 | HL_9DFE_059 | 9DFE | 0.6805 | 1 | GNRA variation | 1A | LSU rRNA | U | 2528 | A | 2530 | A | 2531 | G | 2532 | A | 2533 | A | 2534 | G | 2535 | cWW | ||
| 6 | HL_8P9A_173 | 8P9A | 0.7881 | 1 | GNRA variation | AR | LSU rRNA | A | 2897 | C | 2899 | A | 2900 | G | 2901 | A | 2902 | A | 2903 | U | 2904 | cWW | ||
| 7 | HL_8CRE_061 | 8CRE | 0.7752 | 1 | GNRA variation | 1 | LSU rRNA | A | 2869 | C | 2871 | A | 2872 | G | 2873 | A | 2874 | A | 2875 | U | 2876 | cWW | ||
| 8 | HL_6JXM_002 | 6JXM | 0.6863 | 1 | tRNA anticodon loop | B | tRNA | G | 41 | C | 42 | A | 44 | A | 45 | A | 46 | A | 47 | C | 48 | cWW | ||
| 9 | HL_4LFB_002 | 4LFB | 0.5343 | 0 | A | SSU rRNA | G | 80 | U | 81 | U | 82 | U | 83 | U | 84 | A | 88 | C | 89 | cWW | |||
| 10 | HL_7TZS_001 | 7TZS | 0.4990 | 0 | X | TPP riboswitch (THI element) | C | 27 | U | 28 | G | 29 | C | 30 | G | 31 | U | 32 | G | 33 | cWW | tWW | ||
| 11 | HL_3LA5_002 | 3LA5 | 0.1521 | 1 | Purine riboswitch | A | Purine riboswitch | G | 46 | C | 47 | U | 49 | C | 50 | C | 51 | A | 52 | C | 53 | cWW | ||
| 12 | HL_4LX6_002 | 4LX6 | 0.1037 | 1 | Purine riboswitch | A | Purine riboswitch | G | 46 | C | 47 | U | 49 | C | 50 | C | 51 | A | 52 | C | 53 | cWW | ||
| 13 | HL_4XNR_002 | 4XNR | 0.0000 | 1 | Purine riboswitch | X | Purine riboswitch | G | 46 | U | 47 | U | 49 | C | 50 | U | 51 | A | 52 | C | 53 | cWW | ntWH | |
| 14 | HL_1Y26_002 | 1Y26 | 0.0634 | 1 | Purine riboswitch | X | Purine riboswitch | G | 46 | U | 47 | U | 49 | C | 50 | U | 51 | A | 52 | C | 53 | cWW | ntWH | |
| 15 | HL_1Y27_002 | 1Y27 | 0.0806 | 1 | Purine riboswitch | X | Purine riboswitch | G | 46 | U | 47 | U | 49 | C | 50 | U | 51 | A | 52 | C | 53 | cWW | ||
| 16 | HL_4FEN_002 | 4FEN | 0.0725 | 1 | Purine riboswitch | B | Purine riboswitch | G | 46 | U | 47 | U | 49 | C | 50 | U | 51 | A | 52 | C | 53 | cWW | ntWH | |
| 17 | HL_3RKF_008 | 3RKF | 0.1179 | 1 | Purine riboswitch | C | Purine riboswitch | G | 46 | U | 47 | U | 49 | C | 50 | U | 51 | A | 52 | C | 53 | cWW | ntWH | |
| 18 | HL_9IWF_005 | 9IWF | 0.1442 | 1 | Purine riboswitch | B | Purine riboswitch | G | 32 | U | 33 | U | 35 | C | 36 | G | 37 | A | 38 | C | 39 | cWW | tWW | |
| 19 | HL_5VPO_138 | 5VPO | 0.5399 | 1 | Pseudoknot geometry | XV | P-site ASL SufA6 | U | 32 | U | 33 | C | 34 | G | 35 | G | 36 | 1MG | 37 | A | 39 | ncWW | ||
| 20 | HL_6UFH_003 | 6UFH | 0.5793 | 1 | Pseudoknot geometry | A | RNA (167-MER) | G | 70 | U | 72 | U | 73 | G | 74 | G | 75 | C | 76 | C | 77 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GUUUCUAC | 5 |
| UGAAGAAG | 2 |
| AGCAGAAU | 2 |
| GCUUCCAC | 2 |
| UGCAGAAG | 1 |
| UGUAGUCG | 1 |
| UGAAGUAG | 1 |
| GCUAAAAC | 1 |
| GUUUUAC | 1 |
| CUGCGUG | 1 |
| GUUUCGAC | 1 |
| UUCGGGA | 1 |
| GGUUGGCC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UUUCUA | 5 |
| GCAGAA | 3 |
| GAAGAA | 2 |
| CUUCCA | 2 |
| GUAGUC | 1 |
| GAAGUA | 1 |
| CUAAAA | 1 |
| UUUUA | 1 |
| UGCGU | 1 |
| UUUCGA | 1 |
| UCGG(1MG)G | 1 |
| GUUGGC | 1 |
- Annotations
-
- Purine riboswitch (8)
- GNRA variation (7)
- (2)
- Pseudoknot geometry (2)
- tRNA anticodon loop (1)
- Basepair signature
- cWW-F-F-F-F-F
- Heat map statistics
- Min 0.06 | Avg 0.51 | Max 0.98
Coloring options: