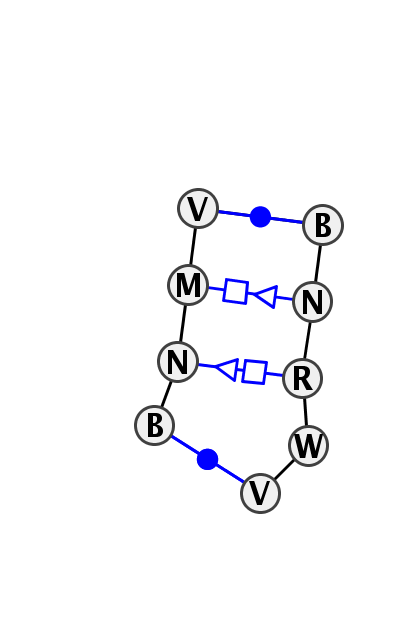
Motif IL_37811.2 Version IL_37811.2 of this group appears in releases 3.8 to 3.11
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | break | 5 | 6 | 7 | 8 | 9 | 1-9 | 2-7 | 2-8 | 3-6 | 4-5 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_3BNR_008 | 3BNR | 0.8074 | 0 | Minor groove platform related | C+D | A site of human mitochondrial ribosome, chain one + A site of human mitochondrial ribosome, chain three | C | 7 | A | 8 | C | 9 | C | 10 | * | G | 13 | C | 14 | A | 15 | A | 16 | G | 17 | cWW | cwW | ncwW | cWW | |
| 2 | IL_3BNT_005 | 3BNT | 0.7374 | 0 | Tandem non-canonical cWW pairs | A | A site of human mitochondrial ribosome | C | 6||||4_665 | A | 7||||4_665 | C | 8||||4_665 | C | 9||||4_665 | * | G | 14 | C | 15 | A | 16 | A | 17 | G | 18 | cWW | cwW | ncBW | cWW | |
| 3 | IL_3BNT_002 | 3BNT | 0.7374 | 0 | Tandem non-canonical cWW pairs | A | A site of human mitochondrial ribosome | C | 6 | A | 7 | C | 8 | C | 9 | * | G | 14||||4_665 | C | 15||||4_665 | A | 16||||4_665 | A | 17||||4_665 | G | 18||||4_665 | cWW | cwW | ncBW | cWW | |
| 4 | IL_3LOA_001 | 3LOA | 0.6669 | 0 | UAA/GAN related | A+B | RNA (21-mer) + RNA (23-mer) | G | 1401 | C | 1402 | C | 1403 | C | 1404 | * | G | 1497 | U | 1498 | A | 1499 | A | 1500 | C | 1501 | cWW | ncWW | cWW | ||
| 5 | IL_4V5O_120 | 4V5O | 0.7753 | 1 | UAA/GAN related | BA | SSU rRNA | U | 728 | U | 729 | A | 730 | C | 731 | * | G | 784 | G | 785 | A | 787 | U | 788 | A | 789 | cWW | ncwW | ncWW | cWW | |
| 6 | IL_3IGI_009 | 3IGI | 0.3003 | 0 | UAA/GAN related | A | Group II catalytic intron D1-D4-3 | U | 172 | G | 173 | A | 174 | G | 175 | * | C | 190 | A | 191 | A | 192 | A | 193 | G | 194 | cWW | tSH | tHS | cWW | |
| 7 | IL_4V8P_183 | 4V8P | 0.2727 | 0 | UAA/GAN variation | D1 | LSU rRNA | U | 571 | G | 572 | A | 573 | G | 574 | * | C | 594 | A | 595 | A | 596 | A | 597 | A | 598 | cWW | ntSH | tHS | cWW | |
| 8 | IL_4V8P_179 | 4V8P | 0.2367 | 0 | UAA/GAN variation | D1 | LSU rRNA | U | 499 | G | 500 | A | 501 | G | 502 | * | C | 475 | A | 476 | G | 477 | A | 478 | G | 479 | cWW | tSH | ntHS | cWW | |
| 9 | IL_2R8S_002 | 2R8S | 0.2868 | 0 | UAA/GAN related | R | P4-P6 RNA RIBOZYME DOMAIN | U | 205 | A | 206 | A | 207 | C | 208 | * | G | 112 | A | 113 | A | 114 | A | 115 | G | 116 | cWW | tSH | tHS | cWW | |
| 10 | IL_5J7L_317 | 5J7L | 0.0000 | 0 | UAA/GAN related | DA | LSU rRNA | U | 1864 | U | 1865 | A | 1866 | G | 1867 | * | C | 1874 | G | 1875 | A | 1876 | A | 1877 | G | 1878 | cWW | tHS | cWW | ||
| 11 | IL_4V8P_278 | 4V8P | 0.2826 | 2 | UAA/GAN variation | D1 | LSU rRNA | U | 3043 | A | 3044 | A | 3046 | A | 3048 | * | U | 3073 | G | 3074 | A | 3075 | A | 3076 | G | 3077 | cWW | tHS | cWW | ||
| 12 | IL_3FTM_002 | 3FTM | 0.3053 | 0 | UAA/GAN related | C+D | RNA (11-mer) + RNA (12-mer) | U | 46 | G | 47 | A | 48 | G | 49 | * | C | -1 | G | 1 | G | 2 | A | 3 | A | 4 | cWW | ntHS | cWW | ||
| 13 | IL_3FS0_001 | 3FS0 | 0.4073 | 0 | UAA/GAN related | A+B | RNA (10-mer) + RNA (11-mer) | U | 46 | G | 47 | A | 48 | G | 49 | * | C | -1 | G | 1 | G | 2 | A | 3 | A | 4 | cWW | nthH | tHS | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CACC*GCAAG | 3 |
| UGAG*CGGAA | 2 |
| GCCC*GUAAC | 1 |
| UUAC*GGAAUA | 1 |
| UGAG*CAAAG | 1 |
| UGAG*CAAAA | 1 |
| UGAG*CAGAG | 1 |
| UAAC*GAAAG | 1 |
| UUAG*CGAAG | 1 |
| UAAAUA*UGAAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AC*CAA | 3 |
| GA*AAA | 2 |
| GA*GGA | 2 |
| CC*UAA | 1 |
| UA*GAAU | 1 |
| GA*AGA | 1 |
| AA*AAA | 1 |
| UA*GAA | 1 |
| AAAU*GAA | 1 |
- Annotations
-
- UAA/GAN related (7)
- UAA/GAN variation (3)
- Tandem non-canonical cWW pairs (2)
- Minor groove platform related (1)
- Basepair signature
- cWW-R-tSH-tHS-cWW
- Heat map statistics
- Min 0.16 | Avg 0.53 | Max 0.92
Coloring options: