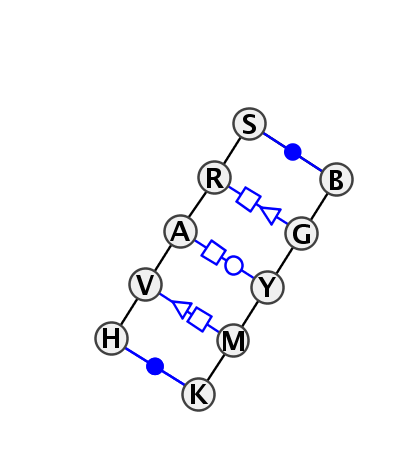
Motif IL_39521.5 Version IL_39521.5 of this group appears in releases 3.54 to 3.76
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | break | 6 | 7 | 8 | 9 | 10 | 1-10 | 2-9 | 3-8 | 4-7 | 5-6 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_2QWY_007 | 2QWY | 0.4868 | 0 | tSH-tWH-tHS | C | SAM-II riboswitch | A | 46 | A | 47 | A | 48 | A | 49 | G | 50 | * | C | 16 | G | 17 | U | 18 | A | 19 | U | 20 | cWW | tSH | tHW | tHS | cWW |
| 2 | IL_5J7L_300 | 5J7L | 0.4475 | 0 | tSH-tWH-tHS | DA | LSU rRNA | U | 1513 | G | 1514 | A | 1515 | G | 1516 | G | 1517 | * | U | 1474 | G | 1475 | U | 1476 | A | 1477 | G | 1478 | cWW | tSH | tHW | tHS | cWW |
| 3 | IL_4V9F_055 | 4V9F | 0.2897 | 0 | tSH-tWH-tHS | 0 | LSU rRNA | U | 1639 | C | 1640 | A | 1641 | A | 1642 | C | 1643 | * | G | 1542 | G | 1543 | U | 1544 | C | 1545 | G | 1546 | cWW | tSH | tHW | tHS | cWW |
| 4 | IL_6CZR_145 | 6CZR | 0.1660 | 0 | tSH-tWH-tHS | 1a | SSU rRNA | C | 763 | A | 764 | A | 765 | A | 766 | C | 767 | * | G | 783 | G | 784 | U | 785 | A | 786 | G | 787 | cWW | tSH | tHW | tHS | cWW |
| 5 | IL_5J7L_037 | 5J7L | 0.1703 | 0 | tSH-tWH-tHS | AA | SSU rRNA | C | 779 | A | 780 | A | 781 | A | 782 | C | 783 | * | G | 799 | G | 800 | U | 801 | A | 802 | G | 803 | cWW | tSH | tHW | tHS | cWW |
| 6 | IL_4LFB_033 | 4LFB | 0.1553 | 0 | tSH-tWH-tHS | A | SSU rRNA | C | 779 | A | 780 | A | 781 | A | 782 | C | 783 | * | G | 799 | G | 800 | U | 801 | A | 802 | G | 803 | cWW | tSH | tHW | tHS | cWW |
| 7 | IL_4V88_436 | 4V88 | 0.0000 | 0 | tSH-tWH-tHS | A6 | SSU rRNA | C | 990 | G | 991 | A | 992 | A | 993 | G | 994 | * | C | 1010 | G | 1011 | U | 1012 | A | 1013 | G | 1014 | cWW | tSH | tHW | tHS | cWW |
| 8 | IL_4V88_418 | 4V88 | 0.1589 | 0 | tSH-tWH-tHS | A6 | SSU rRNA | U | 968 | C | 969 | A | 970 | A | 971 | G | 972 | * | C | 627 | G | 628 | U | 629 | A | 630 | G | 631 | cWW | tSH | tHW | tHS | cWW |
| 9 | IL_6CZR_135 | 6CZR | 0.2094 | 0 | tSH-tWH-tHS | 1a | SSU rRNA | U | 741 | G | 742 | A | 743 | G | 744 | G | 745 | * | U | 564 | G | 565 | U | 566 | A | 567 | G | 568 | cWW | tSH | tHW | ntHS | cWW |
| 10 | IL_4LFB_023 | 4LFB | 0.1872 | 0 | tSH-tWH-tHS | A | SSU rRNA | U | 757 | G | 758 | A | 759 | G | 760 | G | 761 | * | U | 580 | G | 581 | U | 582 | A | 583 | G | 584 | cWW | tSH | tHW | tHS | cWW |
| 11 | IL_5J7L_027 | 5J7L | 0.2117 | 0 | tSH-tWH-tHS | AA | SSU rRNA | U | 757 | C | 758 | A | 759 | G | 760 | G | 761 | * | C | 580 | G | 581 | C | 582 | A | 583 | G | 584 | cWW | tSH | tHW | tHS | cWW |
| 12 | IL_7A0S_049 | 7A0S | 0.2627 | 0 | tSH-tWH-tHS | X | LSU rRNA | U | 1370 | G | 1371 | A | 1372 | G | 1373 | G | 1374 | * | C | 1383 | G | 1384 | C | 1385 | A | 1386 | G | 1387 | cWW | tSH | tHW | tHS | cWW |
| 13 | IL_4V9F_022 | 4V9F | 0.2455 | 0 | tSH-tWH-tHS | 0 | LSU rRNA | C | 719 | G | 720 | A | 721 | G | 722 | G | 723 | * | C | 705 | G | 706 | C | 707 | A | 708 | G | 709 | cWW | tSH | tHW | tHS | cWW |
| 14 | IL_5J7L_293 | 5J7L | 0.2308 | 0 | tSH-tWH-tHS | DA | LSU rRNA | C | 1357 | G | 1358 | A | 1359 | G | 1360 | G | 1361 | * | C | 1370 | G | 1371 | U | 1372 | A | 1373 | G | 1374 | cWW | tSH | tHW | tHS | cWW |
| 15 | IL_7RQB_053 | 7RQB | 0.4421 | 0 | tSH-tWH-tHS | 1A | LSU rRNA | U | 1357 | G | 1358 | A | 1359 | A | 1360 | G | 1361 | * | C | 1370 | G | 1371 | U | 1372 | A | 1373 | G | 1374 | cWW | tSH | ntWW | tHS | cWW |
| 16 | IL_4WF9_052 | 4WF9 | 0.4436 | 0 | tSH-tWH-tHS | X | LSU rRNA | U | 1394 | G | 1395 | A | 1396 | G | 1397 | G | 1398 | * | C | 1407 | G | 1408 | U | 1409 | A | 1410 | G | 1411 | cWW | ntHW | ntHS | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UGAGG*UGUAG | 3 |
| CAAAC*GGUAG | 3 |
| AAAAG*CGUAU | 1 |
| UCAAC*GGUCG | 1 |
| CGAAG*CGUAG | 1 |
| UCAAG*CGUAG | 1 |
| UCAGG*CGCAG | 1 |
| UGAGG*CGCAG | 1 |
| CGAGG*CGCAG | 1 |
| CGAGG*CGUAG | 1 |
| UGAAG*CGUAG | 1 |
| UGAGG*CGUAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAG*GUA | 5 |
| AAA*GUA | 4 |
| GAA*GUA | 2 |
| GAG*GCA | 2 |
| CAA*GUC | 1 |
| CAA*GUA | 1 |
| CAG*GCA | 1 |
Release history
| Release | 3.54 | 3.55 | 3.56 | 3.57 | 3.58 | 3.59 | 3.60 | 3.61 | 3.62 | 3.63 | 3.64 | 3.65 | 3.66 | 3.67 | 3.68 | 3.69 | 3.70 | 3.71 | 3.72 | 3.73 | 3.74 | 3.75 | 3.76 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Date | 2022-02-02 | 2022-03-02 | 2022-03-30 | 2022-04-27 | 2022-05-25 | 2022-06-22 | 2022-07-20 | 2022-08-17 | 2022-10-12 | 2022-11-09 | 2022-12-07 | 2023-01-04 | 2023-02-01 | 2023-03-01 | 2023-03-29 | 2023-04-26 | 2023-05-24 | 2023-06-21 | 2023-07-19 | 2023-08-16 | 2023-09-13 | 2023-10-11 | 2023-11-08 |
| Status | Updated, 1 parent | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- tSH-tWH-tHS (16)
- Basepair signature
- cWW-tSH-tHW-tHS-cWW
- Heat map statistics
- Min 0.06 | Avg 0.33 | Max 0.69
Coloring options: