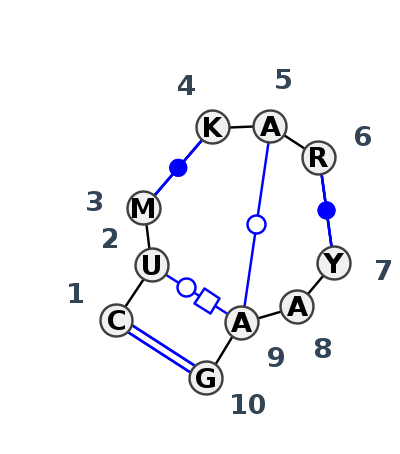
Motif J3_05327.2 Version J3_05327.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | break | 4 | 5 | 6 | break | 7 | 8 | 9 | 10 | 1-10 | 2-9 | 3-4 | 5-9 | 6-7 | 7-9 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | J3_9DFE_016 | 9DFE | 0.2159 | 2 | 1A | LSU rRNA | C | 2295 | U | 2296 | C | 2297 | * | G | 2321 | A | 2322 | G | 2323 | * | U | 2332 | A | 2333 | A | 2335 | G | 2337 | cWW | tWH | cWW | tWW | cWW | ncSs |
| 2 | J3_4WF9_008 | 4WF9 | 0.1650 | 2 | X | LSU rRNA | C | 2322 | U | 2323 | C | 2324 | * | G | 2348 | A | 2349 | G | 2350 | * | C | 2359 | A | 2360 | A | 2362 | G | 2364 | cWW | tWH | cWW | tWW | cWW | |
| 3 | J3_4V9F_008 | 4V9F | 0.1152 | 2 | 0 | LSU rRNA | C | 2329 | U | 2330 | C | 2331 | * | G | 2355 | A | 2356 | G | 2357 | * | C | 2366 | A | 2367 | A | 2369 | G | 2371 | cWW | tWH | cWW | tWW | cWW | |
| 4 | J3_9H3G_023 | 9H3G | 0.0536 | 2 | A | LSU rRNA | C | 2663 | U | 2664 | C | 2665 | * | G | 2689 | A | 2690 | A | 2691 | * | U | 2700 | A | 2701 | A | 2703 | G | 2705 | cWW | tWH | cWW | tWW | cWW | |
| 5 | J3_8GLP_009 | 8GLP | 0.0000 | 2 | L5 | LSU rRNA | C | 4241 | U | 4242 | C | 4243 | * | G | 4267 | A | 4268 | G | 4269 | * | C | 4278 | A | 4279 | A | 4281 | G | 4283 | cWW | tWH | cWW | tWW | cWW | ncSs |
| 6 | J3_9E6Q_016 | 9E6Q | 0.0475 | 2 | 1 | LSU rRNA | C | 2407 | OMU | 2408 | C | 2409 | * | G | 2433 | A | 2434 | G | 2435 | * | C | 2444 | A | 2445 | A | 2447 | G | 2449 | cWW | tWH | cWW | tWW | cWW | |
| 7 | J3_9AXU_022 | 9AXU | 0.0594 | 2 | 2 | LSU rRNA | C | 2759 | U | 2760 | C | 2761 | * | G | 2785 | A | 2786 | A | 2787 | * | U | 2796 | A | 2797 | A | 2799 | G | 2801 | cWW | tWH | cWW | tWW | cWW | |
| 8 | J3_8P9A_062 | 8P9A | 0.0685 | 2 | AR | LSU rRNA | C | 2664 | U | 2665 | C | 2666 | * | G | 2690 | A | 2691 | A | 2692 | * | U | 2701 | A | 2702 | A | 2704 | G | 2706 | cWW | tWH | cWW | tWW | cWW | |
| 9 | J3_8OI5_021 | 8OI5 | 0.0857 | 2 | 1 | LSU rRNA | C | 2636 | U | 2637 | C | 2638 | * | G | 2662 | A | 2663 | A | 2664 | * | U | 2673 | A | 2674 | A | 2676 | G | 2678 | cWW | tWH | cWW | ntWW | cWW | |
| 10 | J3_7A0S_010 | 7A0S | 0.2505 | 2 | X | LSU rRNA | C | 2274 | U | 2275 | C | 2276 | * | G | 2300 | A | 2301 | G | 2302 | * | U | 2311 | A | 2312 | A | 2314 | G | 2316 | cWW | cWW | ncBW | cWW | ||
| 11 | J3_8B0X_022 | 8B0X | 0.2838 | 2 | a | LSU rRNA | C | 2299 | U | 2300 | A | 2301 | * | U | 2325 | A | 2326 | G | 2327 | * | C | 2336 | A | 2337 | A | 2339 | G | 2341 | cWW | tWH | cWW | tWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CUC*GAA*UAAAAG | 4 |
| CUC*GAG*UAGAAG | 2 |
| CUC*GAG*CAAAAG | 2 |
| CUC*GAG*CAUAAG | 1 |
| C(OMU)C*GAG*CAAAAG | 1 |
| CUA*UAG*CAUAAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| U*A*AAAA | 6 |
| U*A*AGAA | 2 |
| U*A*AUAA | 2 |
| (OMU)*A*AAAA | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (11)
- Basepair signature
- cWW-tWH-tWW-cWW-F-cWW
- Heat map statistics
- Min 0.05 | Avg 0.17 | Max 0.35
Coloring options: