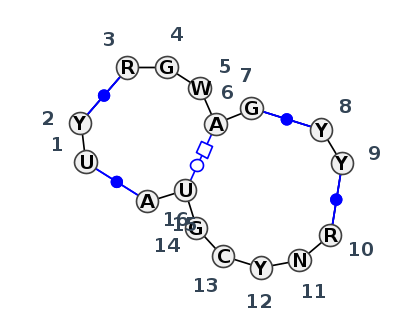
Motif J4_48150.1 Version J4_48150.1 of this group appears in releases 4.0 to 4.0
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | break | 3 | 4 | 5 | 6 | 7 | break | 8 | 9 | break | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 1-16 | 2-3 | 6-15 | 7-8 | 9-10 | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | J4_8B0X_009 | 8B0X | 0.1132 | 1 | a | LSU rRNA | U | 1315 | C | 1316 | * | G | 1340 | G | 1341 | U | 1342 | A | 1344 | G | 1345 | * | C | 1406 | U | 1407 | * | A | 1599 | A | 1600 | U | 1601 | C | 1602 | G | 1603 | U | 1604 | A | 1605 | cWW | cWW | tHW | cWW | cWW |
| 2 | J4_9AXU_005 | 9AXU | 0.0747 | 1 | 2 | LSU rRNA | U | 1529 | C | 1530 | * | G | 1554 | G | 1555 | U | 1556 | A | 1558 | G | 1559 | * | C | 1649 | C | 1650 | * | G | 1884 | G | 1885 | U | 1886 | C | 1887 | G | 1888 | U | 1889 | A | 1890 | cWW | cWW | tHW | cWW | cWW |
| 3 | J4_8GLP_006 | 8GLP | 0.0965 | 1 | L5 | LSU rRNA | U | 2409 | C | 2410 | * | G | 2434 | G | 2435 | U | 2436 | A | 2438 | G | 2439 | * | U | 2538 | C | 2539 | * | G | 2777 | G | 2778 | C | 2779 | C | 2780 | G | 2781 | U | 2782 | A | 2783 | cWW | cWW | tHW | cWW | cWW |
| 4 | J4_4V9F_005 | 4V9F | 0.0763 | 1 | 0 | LSU rRNA | U | 1419 | C | 1420 | * | G | 1444 | G | 1445 | U | 1446 | A | 1448 | G | 1449 | * | C | 1513 | C | 1514 | * | G | 1672 | U | 1673 | C | 1674 | C | 1675 | G | 1676 | U | 1677 | A | 1678 | cWW | cWW | tHW | cWW | cWW |
| 5 | J4_9E6Q_005 | 9E6Q | 0.0000 | 1 | 1 | LSU rRNA | U | 1442 | U | 1443 | * | A | 1467 | G | 1468 | U | 1469 | A | 1471 | G | 1472 | * | C | 1535 | C | 1536 | * | G | 1739 | C | 1740 | C | 1741 | C | 1742 | G | 1743 | U | 1744 | A | 1745 | cWW | cWW | tHW | cWW | cWW |
| 6 | J4_9DFE_005 | 9DFE | 0.1467 | 1 | 1A | LSU rRNA | U | 1313 | C | 1314 | * | G | 1338 | G | 1339 | U | 1340 | A | 1342 | G | 1343 | * | C | 1404 | U | 1405 | * | A | 1597 | C | 1598 | C | 1599 | C | 1600 | G | 1601 | U | 1602 | A | 1603 | cWW | cWW | tHW | cWW | cWW |
| 7 | J4_4WF9_006 | 4WF9 | 0.1660 | 1 | X | LSU rRNA | U | 1350 | C | 1351 | * | G | 1375 | G | 1376 | U | 1377 | A | 1379 | G | 1380 | * | C | 1441 | C | 1442 | * | G | 1641 | C | 1642 | C | 1643 | C | 1644 | G | 1645 | U | 1646 | A | 1647 | cWW | cWW | tHW | cWW | cWW |
| 8 | J4_7A0S_006 | 7A0S | 0.1962 | 1 | X | LSU rRNA | U | 1326 | C | 1327 | * | G | 1351 | G | 1352 | A | 1353 | A | 1355 | G | 1356 | * | C | 1417 | C | 1418 | * | G | 1613 | C | 1614 | C | 1615 | C | 1616 | G | 1617 | U | 1618 | A | 1619 | cWW | cWW | ntHW | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UC*GGUGAG*CU*AAUCGUA | 1 |
| UC*GGUUAG*CC*GGUCGUA | 1 |
| UC*GGUCAG*UC*GGCCGUA | 1 |
| UC*GGUUAG*CC*GUCCGUA | 1 |
| UU*AGUUAG*CC*GCCCGUA | 1 |
| UC*GGUUAG*CU*ACCCGUA | 1 |
| UC*GGUUAG*CC*GCCCGUA | 1 |
| UC*GGAAAG*CC*GCCCGUA | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| *GUUA**CCCGU | 3 |
| *GUGA**AUCGU | 1 |
| *GUUA**GUCGU | 1 |
| *GUCA**GCCGU | 1 |
| *GUUA**UCCGU | 1 |
| *GAAA**CCCGU | 1 |
Release history
| Release | 4.0 |
|---|---|
| Date | 2025-08-13 |
| Status | New id, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (8)
- Basepair signature
- cWW-cWW-tHW-F-F-F-F-F-F-cWW-cWW
- Heat map statistics
- Min 0.07 | Avg 0.13 | Max 0.25
Coloring options: