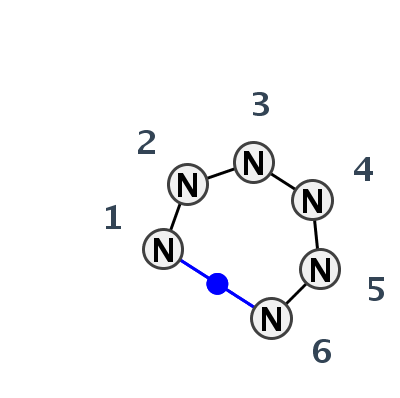
Motif HL_53640.1 Version HL_53640.1 of this group appears in releases 3.77 to 3.79
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 1-6 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_4PQV_002 | 4PQV | 0.6582 | 1 | Pseudoknot geometry | A | Flavivirus 3' UTR stem loop IV | C | 28 | G | 29 | C | 30 | C | 31 | G | 32 | G | 34 | cWW |
| 2 | HL_3V7E_001 | 3V7E | 0.5621 | 1 | Pseudoknot geometry | C | SAM-I riboswitch | G | 23 | C | 25 | U | 26 | G | 27 | G | 28 | C | 29 | cWW |
| 3 | HL_5FJC_001 | 5FJC | 0.5389 | 1 | Pseudoknot geometry | A | SAM-I riboswitch | G | 23 | C | 25 | U | 26 | G | 27 | G | 28 | C | 29 | cWW |
| 4 | HL_4AOB_001 | 4AOB | 0.5721 | 1 | Pseudoknot geometry | A | SAM-I riboswitch | G | 23 | C | 25 | U | 26 | G | 27 | G | 28 | C | 29 | cWW |
| 5 | HL_6MWN_004 | 6MWN | 0.6251 | 1 | Pseudoknot geometry | B | IRES | G | 650 | U | 652 | G | 653 | G | 654 | A | 655 | C | 656 | cWW |
| 6 | HL_4KQY_001 | 4KQY | 0.5771 | 1 | Pseudoknot geometry | A | SAM-I riboswitch | G | 23 | C | 25 | U | 26 | G | 27 | G | 28 | C | 29 | cWW |
| 7 | HL_6VMY_003 | 6VMY | 0.7174 | 1 | Pseudoknot geometry | A | Cobalamin riboswitch | G | 268 | C | 270 | C | 271 | C | 272 | G | 273 | C | 274 | cWW |
| 8 | HL_5U30_002 | 5U30 | 0.7203 | 1 | Pseudoknot geometry | B | sgRNA | U | 43 | U | 45 | C | 46 | C | 47 | A | 48 | G | 49 | cWW |
| 9 | HL_6UFM_005 | 6UFM | 0.6148 | 0 | Pseudoknot geometry | B | RNA (98-MER) | G | 50 | G | 51 | A | 52 | A | 53 | A | 54 | C | 55 | cWW |
| 10 | HL_4WF9_061 | 4WF9 | 0.5891 | 0 | Ribsomal LSU H95 | X | LSU rRNA | U | 2728 | G | 2729 | C | 2730 | C | 2731 | A | 2732 | A | 2733 | cWW |
| 11 | HL_7A0S_064 | 7A0S | 0.6044 | 0 | Ribsomal LSU H95 | X | LSU rRNA | U | 2680 | A | 2681 | C | 2682 | C | 2683 | A | 2684 | A | 2685 | cWW |
| 12 | HL_5J7L_198 | 5J7L | 0.6243 | 0 | Ribsomal LSU H95 | DA | LSU rRNA | U | 2701 | G | 2702 | C | 2703 | C | 2704 | A | 2705 | A | 2706 | cWW |
| 13 | HL_4V9F_067 | 4V9F | 0.6038 | 0 | Ribsomal LSU H95 | 0 | LSU rRNA | C | 2737 | G | 2738 | A | 2739 | G | 2740 | A | 2741 | G | 2742 | cWW |
| 14 | HL_5TBW_066 | 5TBW | 0.5628 | 0 | Ribsomal LSU H95 | 1 | LSU rRNA | U | 3068 | G | 3069 | A | 3070 | U | 3071 | C | 3072 | A | 3073 | cWW |
| 15 | HL_1U6B_002 | 1U6B | 0.5710 | 0 | Pseudoknot geometry | B | 197-MER | C | 70 | G | 71 | C | 72 | C | 73 | C | 74 | G | 75 | cWW |
| 16 | HL_1F27_001 | 1F27 | 0.3982 | 4 | Pseudoknot geometry | A | RNA (19-mer) | U | 7 | G | 12 | G | 13 | A | 14 | C | 15 | A | 16 | cWW |
| 17 | HL_3IGI_005 | 3IGI | 0.2531 | 4 | Pseudoknot geometry | A | Group II catalytic intron D1-D4-3 | U | 178 | A | 183 | A | 184 | C | 185 | A | 186 | A | 187 | cWW |
| 18 | HL_4V88_205 | 4V88 | 0.2547 | 9 | Pseudoknot geometry | A6 | SSU rRNA | A | 829 | U | 839 | U | 840 | U | 841 | C | 842 | U | 843 | cWW |
| 19 | HL_7RQB_014 | 7RQB | 0.0000 | 2 | Pseudoknot geometry (H) | 1A | LSU rRNA | G | 410 | C | 413 | C | 414 | A | 415 | C | 416 | C | 417 | cWW |
| 20 | HL_5J7L_146 | 5J7L | 0.0882 | 2 | Pseudoknot geometry | DA | LSU rRNA | G | 410 | C | 413 | C | 414 | A | 415 | U | 416 | C | 417 | cWW |
| 21 | HL_4WF9_014 | 4WF9 | 0.1320 | 2 | Pseudoknot geometry | X | LSU rRNA | G | 456 | C | 459 | C | 460 | A | 461 | U | 462 | C | 463 | cWW |
| 22 | HL_7A0S_014 | 7A0S | 0.1420 | 2 | Pseudoknot geometry | X | LSU rRNA | G | 423 | C | 426 | C | 427 | A | 428 | C | 429 | C | 430 | cWW |
| 23 | HL_4GXY_003 | 4GXY | 0.2241 | 1 | Pseudoknot geometry | A | Cobalamin riboswitch | G | 65 | C | 67 | C | 68 | C | 69 | G | 70 | C | 71 | cWW |
| 24 | HL_5TBW_060 | 5TBW | 0.2521 | 3 | Pseudoknot geometry | 1 | LSU rRNA | C | 2776 | A | 2780 | U | 2781 | U | 2782 | U | 2783 | G | 2784 | cWW |
| 25 | HL_6N5P_003 | 6N5P | 0.3234 | 3 | Pseudoknot geometry | A | cyclic di-AMP riboswitch | U | 76 | U | 80 | U | 81 | C | 82 | U | 83 | G | 84 | cWW |
| 26 | HL_4WF9_002 | 4WF9 | 0.3532 | 3 | Pseudoknot geometry | X | LSU rRNA | U | 87 | A | 91 | G | 92 | U | 93 | A | 94 | A | 95 | cWW |
| 27 | HL_4V9F_002 | 4V9F | 0.7237 | 3 | Pseudoknot geometry | 0 | LSU rRNA | C | 83 | A | 86 | G | 88 | G | 89 | A | 90 | G | 91 | cWW |
| 28 | HL_7RQB_002 | 7RQB | 0.4601 | 3 | Pseudoknot geometry (H) | 1A | LSU rRNA | C | 87 | A | 92 | G | 93 | C | 94 | G | 94|||A | G | 95 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GACUGGC | 4 |
| GUCCCGC | 2 |
| UGCCAA | 2 |
| GGACCACC | 2 |
| GGACCAUC | 2 |
| CGCCGAG | 1 |
| GCUGGAC | 1 |
| UUUCCAG | 1 |
| GGAAAC | 1 |
| UACCAA | 1 |
| CGAGAG | 1 |
| UGAUCA | 1 |
| CGCCCG | 1 |
| UCAGAGGACA | 1 |
| UGCAUAACAA | 1 |
| AUUUUGUUGGUUUCU | 1 |
| CGGAAUUUG | 1 |
| UACCUUCUG | 1 |
| UGUAAGUAA | 1 |
| CGCACGGAG | 1 |
| CGGUAGCGG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| ACUGG | 4 |
| UCCCG | 2 |
| GCCA | 2 |
| GACCAC | 2 |
| GACCAU | 2 |
| GCCGA | 1 |
| CUGGA | 1 |
| UUCCA | 1 |
| GAAA | 1 |
| ACCA | 1 |
| GAGA | 1 |
| GAUC | 1 |
| GCCC | 1 |
| CAGAGGAC | 1 |
| GCAUAACA | 1 |
| UUUUGUUGGUUUC | 1 |
| GGAAUUU | 1 |
| ACCUUCU | 1 |
| GUAAGUA | 1 |
| GCACGGA | 1 |
| GGUAGCG | 1 |
- Annotations
-
- Pseudoknot geometry (21)
- Ribsomal LSU H95 (5)
- Pseudoknot geometry (H) (2)
- Basepair signature
- cWW-F-F-F-F
- Heat map statistics
- Min 0.09 | Avg 0.50 | Max 0.96
Coloring options: