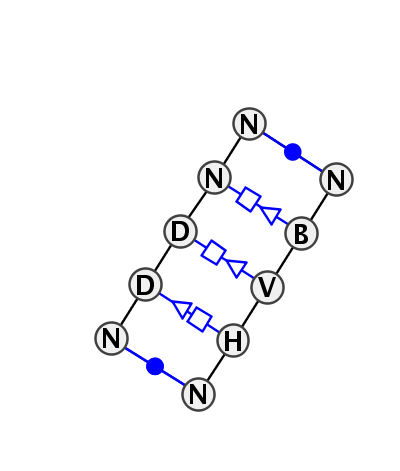
Motif IL_56467.5 Version IL_56467.5 of this group appears in releases 3.48 to 3.53
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | break | 6 | 7 | 8 | 9 | 10 | 1-10 | 2-9 | 2-10 | 3-8 | 4-7 | 5-6 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_7A0S_013 | 7A0S | 0.5904 | 0 | AAA cross-strand stack | X | LSU rRNA | U | 616 | U | 617 | A | 618 | A | 619 | G | 620 | * | C | 629 | G | 630 | G | 631 | A | 632 | G | 633 | cWW | tHS | cWW | |||
| 2 | IL_4WF9_019 | 4WF9 | 0.6284 | 0 | AAA cross-strand stack | X | LSU rRNA | U | 649 | U | 650 | A | 651 | A | 652 | G | 653 | * | U | 663 | G | 664 | G | 665 | A | 666 | G | 667 | cWW | ntWW | tHS | cWW | ||
| 3 | IL_4Y4O_193 | 4Y4O | 0.5394 | 0 | AAA cross-strand stack | 2A | LSU rRNA | U | 606 | U | 607 | A | 608 | A | 609 | G | 610 | * | C | 618 | G | 619 | G | 620 | A | 621 | G | 622 | cWW | ntWH | tHS | cWW | ||
| 4 | IL_5J7L_258 | 5J7L | 0.5278 | 0 | AAA cross-strand stack | DA | LSU rRNA | U | 606 | U | 607 | A | 608 | A | 609 | C | 610 | * | G | 618 | G | 619 | G | 620 | A | 621 | G | 622 | cWW | ntWH | tHS | cWW | ||
| 5 | IL_4V9F_020 | 4V9F | 0.5205 | 0 | AAA cross-strand stack | 0 | LSU rRNA | C | 663 | U | 664 | A | 665 | A | 666 | C | 667 | * | G | 679 | G | 680 | G | 681 | A | 682 | G | 683 | cWW | ntWH | tHS | cWW | ||
| 6 | IL_3MXH_002 | 3MXH | 0.3819 | 0 | Triple sheared | R | Cyclic di-GMP-I riboswitch | C | 22 | A | 23 | A | 24 | A | 25 | C | 26 | * | G | 41 | G | 42 | A | 43 | C | 44 | G | 45 | cWW | ntSH | tHS | cWW | ||
| 7 | IL_4YAZ_006 | 4YAZ | 0.4321 | 0 | Triple sheared | R | Cyclic di-GMP-I riboswitch | U | 16 | A | 17 | A | 18 | A | 19 | C | 20 | * | G | 35 | G | 36 | G | 37 | C | 38 | G | 39 | cWW | ntSH | tHS | cWW | ||
| 8 | IL_3IWN_002 | 3IWN | 0.2994 | 0 | Triple sheared | A | Cyclic di-GMP-I riboswitch | C | 12 | A | 13 | A | 14 | A | 15 | C | 16 | * | G | 31 | G | 32 | A | 33 | C | 34 | G | 35 | cWW | ntSH | tHS | cWW | ||
| 9 | IL_3IGI_002 | 3IGI | 0.2425 | 0 | Triple sheared | A | Group II catalytic intron D1-D4-3 | C | 70 | U | 71 | A | 72 | A | 73 | C | 74 | * | G | 113 | G | 114 | A | 115 | C | 116 | G | 117 | cWW | ntSH | tHS | cWW | ||
| 10 | IL_4Y4O_248 | 4Y4O | 0.0000 | 0 | Triple sheared | 2A | LSU rRNA | C | 1882 | G | 1883 | A | 1884 | A | 1885 | C | 1886 | * | G | 1856 | G | 1857 | G | 1858 | A | 1859 | G | 1860 | cWW | tSH | tHS | tHS | cWW | |
| 11 | IL_5TBW_067 | 5TBW | 0.1982 | 0 | Triple sheared | 1 | LSU rRNA | U | 1645 | G | 1646 | A | 1647 | A | 1648 | U | 1649 | * | A | 1806 | G | 1807 | G | 1808 | A | 1809 | A | 1810 | cWW | tSH | tHS | tHS | cWW | |
| 12 | IL_4WF9_074 | 4WF9 | 0.2874 | 0 | Triple sheared | X | LSU rRNA | C | 1909 | G | 1910 | A | 1911 | A | 1912 | U | 1913 | * | A | 1883 | G | 1884 | G | 1885 | A | 1886 | G | 1887 | ncWW | cSW | tHS | ntHS | cWW | |
| 13 | IL_7A0S_070 | 7A0S | 0.2729 | 0 | Triple sheared | X | LSU rRNA | C | 1865 | G | 1866 | A | 1867 | A | 1868 | A | 1869 | * | U | 1848 | G | 1849 | G | 1850 | A | 1851 | G | 1852 | cWW | ntSH | tHS | cWW | ||
| 14 | IL_2A64_006 | 2A64 | 0.2147 | 0 | Triple sheared | A | RNase P class B | C | 378 | G | 379 | A | 380 | A | 381 | G | 382 | * | C | 347 | G | 348 | G | 349 | A | 350 | G | 351 | cWW | tHS | tHS | cWW | ||
| 15 | IL_2A64_005 | 2A64 | 0.2655 | 0 | Triple sheared | A | RNase P class B | C | 284 | G | 285 | A | 286 | A | 287 | G | 288 | * | C | 297 | G | 298 | G | 299 | A | 300 | G | 301 | cWW | tSH | ntHS | tHS | cWW | |
| 16 | IL_5J7L_310 | 5J7L | 0.2848 | 0 | Triple sheared | DA | LSU rRNA | U | 1720 | G | 1721 | A | 1722 | G | 1723 | G | 1724 | * | U | 1736 | G | 1737 | G | 1738 | A | 1739 | G | 1740 | cWW | tSH | tHS | ntHS | cWW | |
| 17 | IL_4WF9_024 | 4WF9 | 0.3992 | 0 | Triple sheared | X | LSU rRNA | U | 748 | G | 749 | A | 750 | A | 751 | G | 752 | * | U | 769 | G | 770 | G | 771 | A | 772 | G | 773 | cWW | tHS | tHS | cWW | ||
| 18 | IL_4Y4O_198 | 4Y4O | 0.4015 | 0 | Triple sheared | 2A | LSU rRNA | U | 703 | G | 704 | A | 705 | A | 706 | G | 707 | * | U | 724 | G | 725 | G | 726 | A | 727 | G | 728 | cWW | tSH | tHS | tHS | cWW | |
| 19 | IL_5J7L_263 | 5J7L | 0.4072 | 0 | Triple sheared | DA | LSU rRNA | U | 703 | G | 704 | A | 705 | A | 706 | G | 707 | * | U | 724 | G | 725 | G | 726 | A | 727 | G | 728 | cWW | tSH | tHS | tHS | cWW | |
| 20 | IL_5TBW_030 | 5TBW | 0.4364 | 0 | Triple sheared | 1 | LSU rRNA | U | 834 | G | 835 | A | 836 | A | 837 | G | 838 | * | U | 855 | G | 856 | G | 857 | A | 858 | G | 859 | cWW | tSH | tHS | tHS | cWW | |
| 21 | IL_4V9F_027 | 4V9F | 0.3903 | 0 | Triple sheared | 0 | LSU rRNA | U | 794 | G | 795 | A | 796 | A | 797 | G | 798 | * | U | 815 | G | 816 | G | 817 | A | 818 | A | 819 | cWW | ntSH | tHS | tHS | cWW | |
| 22 | IL_4V88_474 | 4V88 | 0.3693 | 0 | Triple sheared | A6 | SSU rRNA | A | 1719 | G | 1720 | A | 1721 | A | 1722 | U | 1723 | * | A | 1678 | G | 1679 | G | 1680 | A | 1681 | U | 1682 | cWW | tHS | tHS | cWW | ||
| 23 | IL_5TBW_106 | 5TBW | 0.4645 | 0 | Triple sheared | 1 | LSU rRNA | C | 2849 | G | 2850 | A | 2851 | C | 2852 | A | 2853 | * | U | 2835 | C | 2836 | A | 2837 | A | 2838 | G | 2839 | cWW | tSH | ntHS | cWW | ||
| 24 | IL_4Y4O_270 | 4Y4O | 0.4181 | 0 | Triple sheared | 2A | LSU rRNA | C | 2480 | G | 2481 | G | 2482 | C | 2483 | G | 2484 | * | C | 2466 | C | 2467 | G | 2468 | A | 2469 | G | 2470 | cWW | tSH | cBW | cWW | ||
| 25 | IL_7A0S_087 | 7A0S | 0.4100 | 0 | Double sheared with non-canonical cWW | X | LSU rRNA | C | 2459 | G | 2460 | G | 2461 | C | 2462 | G | 2463 | * | C | 2445 | C | 2446 | G | 2447 | A | 2448 | G | 2449 | cWW | tSH | ncBW | cWW | ||
| 26 | IL_4WF9_093 | 4WF9 | 0.3441 | 0 | Double sheared with non-canonical cWW | X | LSU rRNA | C | 2507 | G | 2508 | A | 2509 | C | 2510 | G | 2511 | * | C | 2493 | C | 2494 | A | 2495 | A | 2496 | G | 2497 | cWW | ntSH | tHS | cBW | cWW | |
| 27 | IL_5J7L_338 | 5J7L | 0.3381 | 0 | Double sheared with non-canonical cWW | DA | LSU rRNA | C | 2480 | G | 2481 | A | 2482 | C | 2483 | G | 2484 | * | C | 2466 | C | 2467 | A | 2468 | A | 2469 | G | 2470 | cWW | tSH | ntHS | cBW | cWW | |
| 28 | IL_4WFL_001 | 4WFL | 0.3117 | 0 | Double sheared with non-canonical cWW | A | LSRP RNA | C | 91 | G | 92 | A | 93 | C | 94 | A | 95 | * | U | 72 | C | 73 | G | 74 | A | 75 | G | 76 | cWW | tSH | tHS | cBW | cWW | |
| 29 | IL_1U6B_002 | 1U6B | 0.3509 | 0 | Double sheared with non-canonical cWW | B | 197-MER | G | 56 | A | 57 | A | 58 | A | 59 | C | 60 | * | G | 84 | C | 85 | A | 86 | A | 87 | C | 88 | cWW | tSH | tHS | ncWW | cWW | |
| 30 | IL_5J7L_274 | 5J7L | 0.2989 | 0 | Double sheared with non-canonical cWW | DA | LSU rRNA | C | 898 | A | 899 | A | 900 | C | 901 | C | 902 | * | G | 875 | C | 876 | A | 877 | A | 878 | G | 879 | cWW | tSH | ntHS | cBW | cWW | |
| 31 | IL_4V9F_085 | 4V9F | 0.3135 | 0 | Double sheared with non-canonical cWW | 0 | LSU rRNA | C | 2515 | G | 2516 | A | 2517 | C | 2518 | C | 2519 | * | G | 2501 | C | 2502 | A | 2503 | A | 2504 | G | 2505 | cWW | tSH | ntHS | cBW | cWW | |
| 32 | IL_5J7L_056 | 5J7L | 0.4210 | 0 | Double sheared with non-canonical cWW | AA | SSU rRNA | C | 1259 | G | 1260 | A | 1261 | C | 1262 | C | 1263 | * | G | 1272 | C | 1273 | A | 1274 | A | 1275 | G | 1276 | cWW | tSH | tHS | ncBW | cWW | |
| 33 | IL_7A0S_056 | 7A0S | 0.4743 | 0 | Double sheared with near cWW | X | LSU rRNA | C | 1491 | A | 1492 | A | 1493 | G | 1494 | G | 1495 | * | C | 1529 | U | 1530 | C | 1531 | A | 1532 | G | 1533 | ncWW | tSH | cWW | |||
| 34 | IL_4WF9_084 | 4WF9 | 0.7665 | 0 | Other IL | X | LSU rRNA | A | 2122 | A | 2123 | U | 2124 | U | 2125 | C | 2126 | * | G | 2217 | G | 2218 | C | 2219 | U | 2220 | U | 2221 | cWW | ncWW | ncWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UGAAG*UGGAG | 4 |
| UUAAG*CGGAG | 2 |
| CAAAC*GGACG | 2 |
| CGAAG*CGGAG | 2 |
| CGGCG*CCGAG | 2 |
| CGACG*CCAAG | 2 |
| CGACC*GCAAG | 2 |
| UUAAG*UGGAG | 1 |
| UUAAC*GGGAG | 1 |
| CUAAC*GGGAG | 1 |
| UAAAC*GGGCG | 1 |
| CUAAC*GGACG | 1 |
| CGAAC*GGGAG | 1 |
| UGAAU*AGGAA | 1 |
| CGAAU*AGGAG | 1 |
| CGAAA*UGGAG | 1 |
| UGAGG*UGGAG | 1 |
| UGAAG*UGGAA | 1 |
| AGAAU*AGGAU | 1 |
| CGACA*UCAAG | 1 |
| CGACA*UCGAG | 1 |
| GAAAC*GCAAC | 1 |
| CAACC*GCAAG | 1 |
| CAAGG*CUCAG | 1 |
| AAUUC*GGCUU | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAA*GGA | 12 |
| UAA*GGA | 5 |
| GAC*CAA | 5 |
| AAA*GAC | 2 |
| GGC*CGA | 2 |
| AAA*GGC | 1 |
| UAA*GAC | 1 |
| GAG*GGA | 1 |
| GAC*CGA | 1 |
| AAA*CAA | 1 |
| AAC*CAA | 1 |
| AAG*UCA | 1 |
| AUU*GCU | 1 |
- Annotations
-
- Triple sheared (19)
- Double sheared with non-canonical cWW (8)
- AAA cross-strand stack (5)
- Other IL (1)
- Double sheared with near cWW (1)
- Basepair signature
- cWW-tSH-tHS-tHS-cWW
- Heat map statistics
- Min 0.07 | Avg 0.47 | Max 0.98
Coloring options: