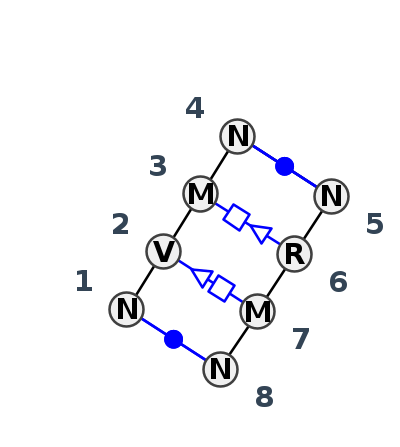
Motif IL_58355.3 Version IL_58355.3 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | break | 5 | 6 | 7 | 8 | 1-8 | 2-6 | 2-7 | 3-6 | 4-5 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_8GLP_190 | 8GLP | 0.3594 | 1 | S2 | SSU rRNA | U | 514 | G | 515 | OMC | 517 | G | 518 | * | C | 37 | A | 38 | A | 39 | A | 40 | cWW | tSH | tHS | cWW | ||
| 2 | IL_9H3G_127 | 9H3G | 0.2902 | 1 | A | LSU rRNA | C | 3002 | G | 3003 | A | 3004 | U | 3005 | * | A | 3137 | G | 3138 | A | 3139 | G | 3141 | cWW | tSH | tHS | cWW | ||
| 3 | IL_9AXU_107 | 9AXU | 0.3605 | 2 | Double sheared (H) | 2 | LSU rRNA | C | 3100 | A | 3101 | A | 3102 | U | 3103 | * | A | 3235 | G | 3236 | A | 3237 | G | 3240 | cWW | tSH | tHS | cWW | |
| 4 | IL_8P9A_342 | 8P9A | 0.3876 | 2 | Double sheared | AR | LSU rRNA | A | 3139 | G | 3140 | A | 3141 | G | 3144 | * | C | 3004 | A | 3005 | A | 3006 | U | 3007 | cWW | tSH | tHS | cWW | |
| 5 | IL_8OI5_114 | 8OI5 | 0.3799 | 2 | Double sheared (H) | 1 | LSU rRNA | A | 3111 | G | 3112 | A | 3113 | G | 3116 | * | C | 2976 | A | 2977 | A | 2978 | U | 2979 | cWW | tSH | tHS | cWW | |
| 6 | IL_4LFB_067 | 4LFB | 0.3303 | 0 | Double sheared | A | SSU rRNA | C | 1259 | C | 1260 | A | 1261 | C | 1262 | * | G | 1273 | G | 1274 | A | 1275 | G | 1276 | cWW | ncWW | ntSH | tHS | cWW |
| 7 | IL_4UYK_005 | 4UYK | 0.2406 | 0 | Double sheared | R | SRP RNA | C | 121 | C | 122 | A | 123 | C | 124 | * | G | 77 | G | 78 | C | 79 | G | 80 | cWW | ncWw | tHS | cWW | |
| 8 | IL_4ZNP_002 | 4ZNP | 0.2137 | 0 | Double sheared | A | ZMP/ZTP riboswitch | G | 9 | G | 10 | A | 11 | C | 12 | * | G | 37 | G | 38 | A | 39 | C | 40 | cWW | tSH | tHS | cWW | |
| 9 | IL_8P9A_460 | 8P9A | 0.2650 | 0 | Double sheared | sR | SSU rRNA | U | 1669 | G | 1670 | A | 1671 | G | 1672 | * | C | 1729 | A | 1730 | A | 1731 | A | 1732 | cWW | tSH | tHS | cWW | |
| 10 | IL_8GLP_282 | 8GLP | 0.2237 | 0 | Double sheared (H) | S2 | SSU rRNA | U | 1733 | G | 1734 | A | 1735 | G | 1736 | * | C | 1798 | G | 1799 | A | 1800 | A | 1801 | cWW | tSH | tHS | cWW | |
| 11 | IL_8CRE_474 | 8CRE | 0.2467 | 0 | Double sheared | CM | SSU rRNA | U | 1656 | G | 1657 | A | 1658 | G | 1659 | * | C | 1716 | A | 1717 | A | 1718 | A | 1719 | cWW | tSH | tHS | cWW | |
| 12 | IL_9H3G_241 | 9H3G | 0.2113 | 0 | h1 | SSU rRNA | U | 1671 | G | 1672 | A | 1673 | A | 1674 | * | U | 1733 | A | 1734 | A | 1735 | A | 1736 | cWW | tSH | tHS | cWW | ||
| 13 | IL_8B0X_076 | 8B0X | 0.2136 | 0 | Double sheared (H) | a | LSU rRNA | U | 556 | G | 557 | A | 558 | C | 559 | * | G | 536 | G | 537 | A | 538 | G | 539 | cWW | tSH | tHS | cWW | |
| 14 | IL_4WF9_016 | 4WF9 | 0.1640 | 0 | Double sheared | X | LSU rRNA | U | 597 | G | 598 | A | 599 | U | 600 | * | A | 581 | G | 582 | A | 583 | G | 584 | cWW | tSH | tHS | cWW | |
| 15 | IL_9DFE_089 | 9DFE | 0.0967 | 0 | Double sheared | 1A | LSU rRNA | C | 2350 | G | 2351 | A | 2352 | G | 2353 | * | C | 2364 | G | 2365 | A | 2366 | G | 2367 | cWW | tSH | tHS | cWW | |
| 16 | IL_8B0X_148 | 8B0X | 0.0000 | 0 | Double sheared (H) | a | LSU rRNA | C | 2354 | G | 2355 | A | 2356 | G | 2357 | * | C | 2368 | G | 2369 | A | 2370 | G | 2371 | cWW | tSH | tHS | cWW | |
| 17 | IL_4WF9_087 | 4WF9 | 0.1120 | 0 | Double sheared | X | LSU rRNA | C | 2377 | G | 2378 | A | 2379 | G | 2380 | * | C | 2391 | G | 2392 | A | 2393 | G | 2394 | cWW | tSH | tHS | cWW | |
| 18 | IL_7A0S_081 | 7A0S | 0.1442 | 0 | Double sheared | X | LSU rRNA | C | 2329 | G | 2330 | A | 2331 | G | 2332 | * | C | 2343 | G | 2344 | A | 2345 | G | 2346 | cWW | tSH | ntHS | cWW | |
| 19 | IL_8CRE_402 | 8CRE | 0.1684 | 0 | Double sheared | CM | SSU rRNA | C | 184 | G | 185 | A | 186 | C | 187 | * | G | 194 | A | 195 | A | 196 | G | 197 | cWW | tSH | tHS | cWW | |
| 20 | IL_8GLP_052 | 8GLP | 0.1548 | 0 | L5 | LSU rRNA | C | 1450 | G | 1451 | A | 1452 | G | 1453 | * | C | 1431 | G | 1432 | A | 1433 | G | 1434 | cWW | tSH | tHS | cWW | ||
| 21 | IL_1SAQ_005 | 1SAQ | 0.1749 | 0 | Double sheared | C+D | RNA (8-mer) | C | 203 | G | 204 | A | 205 | G | 206 | * | C | 211 | G | 212 | A | 213 | G | 214 | cWW | ntSH | tHS | cWW | |
| 22 | IL_1SA9_002 | 1SA9 | 0.1580 | 0 | Double sheared | D+C | RNA (8-mer) | C | 211 | G | 212 | A | 213 | G | 214 | * | C | 203 | G | 204 | A | 205 | G | 206 | cWW | ntSH | tHS | cWW | |
| 23 | IL_3D0U_002 | 3D0U | 0.1809 | 0 | Double sheared | A | Lysine riboswitch | C | 67 | G | 68 | A | 69 | G | 70 | * | C | 16 | G | 17 | A | 18 | G | 19 | cWW | tSH | tHS | cWW | |
| 24 | IL_5BTP_004 | 5BTP | 0.1764 | 0 | Double sheared | B | ZMP/ZTP riboswitch | U | 12 | G | 13 | A | 14 | C | 15 | * | G | 41 | A | 42 | A | 43 | G | 44 | cWW | tSH | tHS | cWW | |
| 25 | IL_9BZC_001 | 9BZC | 0.1972 | 0 | Double sheared | A | RNA (87-MER) | U | 9 | G | 10 | A | 11 | C | 12 | * | G | 36 | A | 37 | A | 38 | G | 39 | cWW | tSH | tHS | cWW | |
| 26 | IL_4XWF_001 | 4XWF | 0.1689 | 0 | Double sheared | A | ZMP/ZTP riboswitch | U | 7 | G | 8 | A | 9 | C | 10 | * | G | 36 | G | 37 | A | 38 | G | 39 | cWW | tSH | tHS | cWW | |
| 27 | IL_4LFB_060 | 4LFB | 0.1720 | 0 | Double sheared | A | SSU rRNA | U | 1481 | G | 1482 | A | 1483 | C | 1484 | * | G | 1416 | G | 1417 | A | 1418 | G | 1419 | cWW | tSH | tHS | cWW | |
| 28 | IL_4KZD_002 | 4KZD | 0.2070 | 0 | Double sheared | R | RNA (84-MER) | U | 71 | A | 72 | A | 73 | C | 74 | * | G | 11 | A | 12 | A | 13 | A | 14 | cWW | tSH | tHS | cWW | |
| 29 | IL_5AOX_003 | 5AOX | 0.1486 | 0 | Double sheared | C | ALU JO CONSENSUS RNA | A | 105 | G | 106 | A | 107 | C | 108 | * | G | 58 | G | 59 | A | 60 | U | 61 | cWW | tSH | tHS | cWW | |
| 30 | IL_8P9A_385 | 8P9A | 0.2301 | 0 | Double sheared | sR | SSU rRNA | C | 186 | G | 187 | A | 188 | C | 189 | * | G | 196 | A | 197 | A | 198 | G | 199 | cWW | tSH | tHS | cWW | |
| 31 | IL_6YL5_010 | 6YL5 | 0.2544 | 0 | Double sheared | I | Chains: A,B,C,D,E,F,G,H,I,J,K,L | C | 12 | A | 13 | A | 14 | C | 15 | * | G | 24 | G | 25 | C | 26 | G | 27 | cWW | tSH | tHS | cWW | |
| 32 | IL_6YML_002 | 6YML | 0.2670 | 0 | Double sheared | A | Chains: A,C | C | 12 | A | 13 | A | 14 | C | 15 | * | G | 24 | G | 25 | C | 26 | G | 27 | cWW | tSH | ntHS | cWW | |
| 33 | IL_8T29_001 | 8T29 | 0.2877 | 0 | Double sheared; A in syn | R | RNA (90-MER) | G | 80 | A | 81 | A | 82 | G | 83 | * | C | 8 | G | 9 | A | 10 | C | 11 | cWW | tSH | ntWS | cWW | |
| 34 | IL_9H3G_238 | 9H3G | 0.2762 | 0 | h1 | SSU rRNA | C | 1655 | G | 1656 | A | 1657 | U | 1658 | * | A | 1748 | G | 1749 | A | 1750 | G | 1751 | cWW | tSH | tHS | cWW | ||
| 35 | IL_8P9A_457 | 8P9A | 0.2982 | 0 | Double sheared | sR | SSU rRNA | C | 1653 | G | 1654 | A | 1655 | U | 1656 | * | A | 1744 | G | 1745 | A | 1746 | G | 1747 | cWW | tSH | tHS | cWW | |
| 36 | IL_8GLP_279 | 8GLP | 0.3050 | 0 | Double sheared (H) | S2 | SSU rRNA | C | 1717 | G | 1718 | A | 1719 | U | 1720 | * | A | 1813 | G | 1814 | A | 1815 | G | 1816 | cWW | tSH | tHS | cWW | |
| 37 | IL_8CRE_471 | 8CRE | 0.2739 | 0 | Double sheared | CM | SSU rRNA | C | 1640 | G | 1641 | A | 1642 | U | 1643 | * | A | 1731 | G | 1732 | A | 1733 | G | 1734 | cWW | tSH | tHS | cWW | |
| 38 | IL_8B0X_061 | 8B0X | 0.4562 | 0 | Double sheared (H) | A | SSU rRNA | G | 1416 | G | 1417 | A | 1418 | G | 1419 | * | U | 1481 | G | 1482 | A | 1483 | C | 1484 | cWW | tSH | tHS | cWW | |
| 39 | IL_8OI5_132 | 8OI5 | 0.4177 | 0 | Double sheared; A in syn (H) | 1 | LSU rRNA | A | 3307 | G | 3308 | A | 3309 | G | 3310 | * | C | 3325 | G | 3326 | A | 3327 | U | 3328 | cWW | cWW | |||
| 40 | IL_8P9A_362 | 8P9A | 0.4484 | 0 | Double sheared; A in syn | AR | LSU rRNA | A | 3342 | G | 3343 | A | 3344 | G | 3345 | * | C | 3360 | G | 3361 | A | 3362 | U | 3363 | cWW | cWW | |||
| 41 | IL_9AXU_123 | 9AXU | 0.4282 | 0 | Double sheared; A in syn (H) | 2 | LSU rRNA | A | 3443 | G | 3444 | A | 3445 | G | 3446 | * | C | 3461 | G | 3462 | A | 3463 | U | 3464 | cWW | tHS | cWW | ||
| 42 | IL_4V9F_100 | 4V9F | 0.3707 | 0 | Double sheared | 0 | LSU rRNA | C | 2873 | G | 2874 | A | 2875 | G | 2876 | * | C | 2881 | G | 2882 | A | 2883 | G | 2884 | cWW | tSH | tHS | cWW | |
| 43 | IL_8GLP_179 | 8GLP | 0.3342 | 0 | L5 | LSU rRNA | A | 5014 | G | 5015 | A | 5016 | G | 5017 | * | C | 5032 | G | 5033 | A | 5034 | U | 5035 | cWW | tSH | tHS | cWW | ||
| 44 | IL_9H3G_146 | 9H3G | 0.3563 | 0 | A | LSU rRNA | A | 3329 | G | 3330 | A | 3331 | G | 3332 | * | C | 3347 | G | 3348 | A | 3349 | U | 3350 | cWW | tSH | tHS | cWW | ||
| 45 | IL_9E6Q_104 | 9E6Q | 0.3733 | 0 | Double sheared (H) | 1 | LSU rRNA | C | 2975 | G | 2976 | A | 2977 | G | 2978 | * | C | 2986 | G | 2987 | A | 2988 | G | 2989 | cWW | tSH | tHS | cWW | |
| 46 | IL_9E6Q_100 | 9E6Q | 0.4136 | 0 | 1 | LSU rRNA | C | 3016 | G | 3017 | A | 3018 | G | 3019 | * | C | 2927 | G | 2928 | A | 2929 | G | 2930 | cWW | tSH | tHS | cWW | ||
| 47 | IL_3DIL_002 | 3DIL | 0.4909 | 0 | Double sheared | A | Lysine riboswitch | C | 70 | G | 71 | A | 72 | G | 73 | * | C | 19 | G | 20 | A | 21 | G | 22 | cWW | tSH | tHS | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CGAG*CGAG | 12 |
| AGAG*CGAU | 5 |
| CGAU*AGAG | 4 |
| UGAC*GGAG | 3 |
| AGAACG*CAAU | 2 |
| UGAG*CAAA | 2 |
| CGAC*GAAG | 2 |
| UGAC*GAAG | 2 |
| CAAC*GGCG | 2 |
| UGA(OMC)G*CAAA | 1 |
| CGAU*AGAAG | 1 |
| CAAU*AGAAAG | 1 |
| CCAC*GGAG | 1 |
| CCAC*GGCG | 1 |
| GGAC*GGAC | 1 |
| UGAG*CGAA | 1 |
| UGAA*UAAA | 1 |
| UGAU*AGAG | 1 |
| UAAC*GAAA | 1 |
| AGAC*GGAU | 1 |
| GAAG*CGAC | 1 |
| GGAG*UGAC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GA*GA | 29 |
| GA*AA | 7 |
| GAAC*AA | 2 |
| AA*GC | 2 |
| GA(OMC)*AA | 1 |
| GA*GAA | 1 |
| AA*GAAA | 1 |
| CA*GA | 1 |
| CA*GC | 1 |
| AA*AA | 1 |
| AA*GA | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- Double sheared (27)
- Double sheared (H) (8)
- (8)
- Double sheared; A in syn (H) (2)
- Double sheared; A in syn (2)
- Basepair signature
- cWW-tSH-tHS-cWW
- Heat map statistics
- Min 0.05 | Avg 0.34 | Max 0.84
Coloring options: