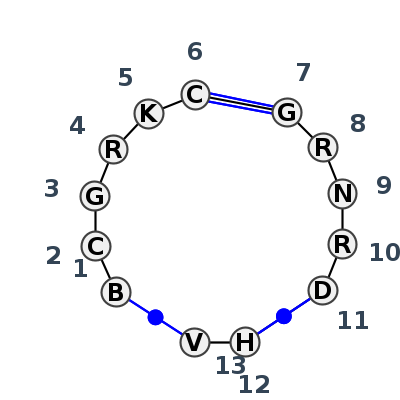
Motif J3_94309.2 Version J3_94309.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | break | 7 | 8 | 9 | 10 | 11 | break | 12 | 13 | 1-13 | 5-8 | 6-7 | 11-12 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | J3_8GLP_030 | 8GLP | 0.1469 | 0 | L5 | LSU rRNA | U | 2440 | C | 2441 | G | 2442 | G | 2443 | U | 2444 | C | 2445 | * | G | 2515 | G | 2516 | A | 2517 | G | 2518 | U | 2519 | * | A | 2536 | A | 2537 | cWW | ntHS | cWW | cWW |
| 2 | J3_4V9F_016 | 4V9F | 0.0989 | 0 | 0 | LSU rRNA | C | 1450 | C | 1451 | G | 1452 | G | 1453 | U | 1454 | C | 1455 | * | G | 1490 | G | 1491 | A | 1492 | A | 1493 | A | 1494 | * | U | 1511 | G | 1512 | cWW | ntHS | cWW | cWW |
| 3 | J3_8OI5_014 | 8OI5 | 0.0000 | 0 | 1 | LSU rRNA | U | 1522 | C | 1523 | G | 1524 | A | 1525 | U | 1526 | C | 1527 | * | G | 1587 | G | 1588 | A | 1589 | A | 1590 | U | 1591 | * | A | 1608 | A | 1609 | cWW | ntHS | cWW | cWW |
| 4 | J3_8P9A_055 | 8P9A | 0.0729 | 0 | AR | LSU rRNA | U | 1526 | C | 1527 | G | 1528 | A | 1529 | U | 1530 | C | 1531 | * | G | 1591 | G | 1592 | A | 1593 | A | 1594 | U | 1595 | * | A | 1612 | A | 1613 | cWW | ntHS | cWW | cWW |
| 5 | J3_9AXU_015 | 9AXU | 0.0746 | 0 | 2 | LSU rRNA | U | 1560 | C | 1561 | G | 1562 | A | 1563 | U | 1564 | C | 1565 | * | G | 1626 | G | 1627 | A | 1628 | A | 1629 | U | 1630 | * | A | 1647 | A | 1648 | cWW | ntHS | cWW | cWW |
| 6 | J3_9H3G_017 | 9H3G | 0.0953 | 0 | A | LSU rRNA | U | 1535 | C | 1536 | G | 1537 | A | 1538 | U | 1539 | C | 1540 | * | G | 1592 | G | 1593 | G | 1594 | A | 1595 | U | 1596 | * | A | 1613 | A | 1614 | cWW | ntHS | cWW | cWW |
| 7 | J3_9E6Q_011 | 9E6Q | 0.1034 | 0 | 1 | LSU rRNA | C | 1473 | C | 1474 | G | 1475 | G | 1476 | U | 1477 | C | 1478 | * | G | 1512 | G | 1513 | A | 1514 | A | 1515 | A | 1516 | * | U | 1533 | G | 1534 | cWW | ntHS | cWW | cWW |
| 8 | J3_9DFE_012 | 9DFE | 0.4929 | 1 | 1A | LSU rRNA | G | 1344 | C | 1345 | G | 1346 | G | 1347 | G | 1348 | C | 1350 | * | G | 1381 | G | 1382 | C | 1383 | A | 1384 | G | 1385 | * | C | 1402 | C | 1403 | cWW | cWW | cWW | |
| 9 | J3_7A0S_018 | 7A0S | 0.5129 | 1 | X | LSU rRNA | U | 1357 | C | 1358 | G | 1359 | G | 1360 | G | 1361 | C | 1363 | * | G | 1394 | A | 1395 | C | 1396 | A | 1397 | G | 1398 | * | C | 1415 | A | 1416 | cWW | cWW | cWW | |
| 10 | J3_4WF9_016 | 4WF9 | 0.5577 | 1 | X | LSU rRNA | U | 1381 | C | 1382 | G | 1383 | G | 1384 | G | 1385 | C | 1387 | * | G | 1418 | A | 1419 | U | 1420 | A | 1421 | A | 1422 | * | U | 1439 | A | 1440 | cWW | cWW | ncWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UCGAUC*GGAAU*AA | 3 |
| CCGGUC*GGAAA*UG | 2 |
| UCGGUC*GGAGU*AA | 1 |
| UCGAUC*GGGAU*AA | 1 |
| GCGGGAC*GGCAG*CC | 1 |
| UCGGGAC*GACAG*CA | 1 |
| UCGGGUC*GAUAA*UA | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| CGAU*GAA* | 3 |
| CGGU*GAA* | 2 |
| CGGU*GAG* | 1 |
| CGAU*GGA* | 1 |
| CGGGA*GCA* | 1 |
| CGGGA*ACA* | 1 |
| CGGGU*AUA* | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (10)
- Basepair signature
- cWW-F-cWW-F-F-F-F-F-cWW-F
- Heat map statistics
- Min 0.07 | Avg 0.28 | Max 0.58
Coloring options: