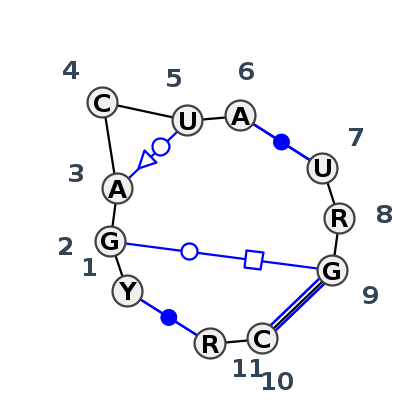
Motif J3_46658.2 Version J3_46658.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | break | 7 | 8 | 9 | break | 10 | 11 | 1-11 | 2-9 | 3-5 | 6-7 | 9-10 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | J3_9DFE_007 | 9DFE | 0.1999 | 3 | 1A | LSU rRNA | C | 683 | G | 684 | A | 685 | C | 687 | U | 688 | A | 689 | * | U | 773 | A | 774 | G | 775 | * | C | 791 | G | 794 | cWW | tWH | tSW | cWW | cWW |
| 2 | J3_7A0S_004 | 7A0S | 0.1296 | 3 | X | LSU rRNA | U | 696 | G | 697 | A | 698 | C | 700 | U | 701 | A | 702 | * | U | 786 | A | 787 | G | 788 | * | C | 804 | A | 807 | cWW | tWH | tSW | cWW | cWW |
| 3 | J3_4WF9_002 | 4WF9 | 0.1008 | 3 | X | LSU rRNA | U | 728 | G | 729 | A | 730 | C | 732 | U | 733 | A | 734 | * | U | 818 | A | 819 | G | 820 | * | C | 836 | A | 839 | cWW | tWH | tSW | cWW | cWW |
| 4 | J3_4V9F_002 | 4V9F | 0.0871 | 3 | 0 | LSU rRNA | C | 774 | G | 775 | A | 776 | C | 778 | U | 779 | A | 780 | * | U | 866 | A | 867 | G | 868 | * | C | 884 | G | 887 | cWW | tWH | tSW | cWW | cWW |
| 5 | J3_8B0X_016 | 8B0X | 0.0742 | 3 | a | LSU rRNA | U | 685 | G | 686 | A | 687 | C | 689 | U | 690 | A | 691 | * | U | 775 | G | 776 | G | 777 | * | C | 793 | A | 796 | cWW | tWH | tSW | cWW | cWW |
| 6 | J3_9E6Q_007 | 9E6Q | 0.0630 | 3 | 1 | LSU rRNA | C | 800 | G | 801 | A | 802 | C | 804 | U | 805 | A | 806 | * | U | 891 | A | 892 | G | 893 | * | C | 909 | G | 912 | cWW | tWH | tSW | cWW | cWW |
| 7 | J3_9H3G_012 | 9H3G | 0.0691 | 3 | A | LSU rRNA | U | 823 | G | 824 | A | 825 | C | 827 | PSU | 828 | A | 829 | * | U | 914 | A | 915 | G | 916 | * | C | 932 | A | 935 | cWW | tWH | tSW | cWW | cWW |
| 8 | J3_9AXU_009 | 9AXU | 0.0512 | 3 | 2 | LSU rRNA | U | 846 | G | 847 | A | 848 | C | 850 | U | 851 | A | 852 | * | U | 937 | A | 938 | G | 939 | * | C | 955 | A | 958 | cWW | tWH | tSW | cWW | cWW |
| 9 | J3_8P9A_049 | 8P9A | 0.0521 | 3 | AR | LSU rRNA | U | 814 | G | 815 | A | 816 | C | 818 | U | 819 | A | 820 | * | U | 905 | A | 906 | G | 907 | * | C | 923 | A | 926 | cWW | tWH | tSW | cWW | cWW |
| 10 | J3_8OI5_009 | 8OI5 | 0.0000 | 3 | 1 | LSU rRNA | U | 810 | G | 811 | A | 812 | C | 814 | U | 815 | A | 816 | * | U | 901 | A | 902 | G | 903 | * | C | 919 | A | 922 | cWW | tWH | tSW | cWW | cWW |
| 11 | J3_8GLP_002 | 8GLP | 0.1652 | 3 | L5 | LSU rRNA | U | 1531 | G | 1532 | A | 1533 | C | 1535 | PSU | 1536 | A | 1537 | * | U | 1622 | A | 1623 | G | 1624 | * | C | 1640 | A | 1643 | cWW | tWH | tSW | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UGAACUA*UAG*CGAA | 3 |
| UGA(A2M)C(PSU)A*UAG*CGAA | 2 |
| CGAGCUA*UAG*CGAG | 1 |
| UGAGCUA*UAG*CGAA | 1 |
| UGAUCUA*UAG*CGAA | 1 |
| CGAUCUA*UAG*CGAG | 1 |
| UGAUCUA*UGG*CAAA | 1 |
| CGAUCUA*UAG*CAAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAACU*A*GA | 3 |
| GAGCU*A*GA | 2 |
| GAUCU*A*GA | 2 |
| GA(A2M)C(PSU)*A*GA | 2 |
| GAUCU*G*AA | 1 |
| GAUCU*A*AA | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (11)
- Basepair signature
- cWW-tWH-cWW-tSW-F-F-cWW
- Heat map statistics
- Min 0.05 | Avg 0.12 | Max 0.36
Coloring options: