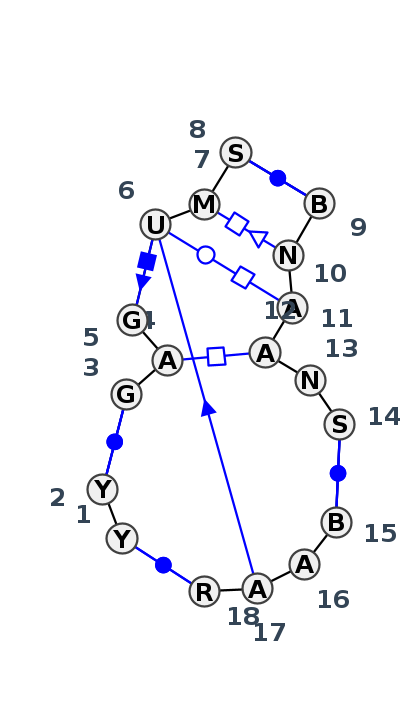
Motif J4_94698.2 Version J4_94698.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | break | 3 | 4 | 5 | 6 | 7 | 8 | break | 9 | 10 | 11 | 12 | 13 | 14 | break | 15 | 16 | 17 | 18 | 1-18 | 2-3 | 4-12 | 5-6 | 6-11 | 6-17 | 7-10 | 8-9 | 11-16 | 12-14 | 12-16 | 12-17 | 14-15 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | J4_9H3G_001 | 9H3G | 0.1112 | 1 | A | LSU rRNA | C | 110 | C | 111 | * | G | 264 | A | 265 | G | 266 | U | 267 | C | 268 | G | 269 | * | C | 290 | C | 291 | A | 292 | A | 293 | U | 294 | G | 296 | * | C | 313 | A | 314 | A | 315 | G | 316 | cWW | cWW | tHH | cSH | tWH | cSs | tHS | cWW | ntWW | ntWW | ntWW | cWW | |
| 2 | J4_8P9A_012 | 8P9A | 0.0778 | 1 | AR | LSU rRNA | U | 112 | C | 113 | * | G | 267 | A | 268 | G | 269 | U | 270 | C | 271 | G | 272 | * | C | 293 | U | 294 | A | 295 | A | 296 | G | 297 | G | 299 | * | U | 316 | A | 317 | A | 318 | A | 319 | cWW | cWW | tHH | cSH | tWH | ncSs | tHS | cWW | ntWW | ntWW | cWW | ||
| 3 | J4_8GLP_001 | 8GLP | 0.0674 | 1 | L5 | LSU rRNA | C | 111 | C | 112 | * | G | 278 | A | 279 | G | 280 | U | 281 | C | 282 | G | 283 | * | C | 304 | A | 305 | A | 306 | A | 307 | G | 308 | G | 310 | * | U | 327 | A | 328 | A | 329 | G | 330 | cWW | cWW | tHH | cSH | tWH | cSs | tHS | cWW | ntWW | ntWW | ntWW | cWW | |
| 4 | J4_8OI5_001 | 8OI5 | 0.0000 | 1 | 1 | LSU rRNA | U | 111 | C | 112 | * | G | 267 | A | 268 | G | 269 | U | 270 | C | 271 | G | 272 | * | C | 293 | U | 294 | A | 295 | A | 296 | G | 297 | G | 299 | * | U | 316 | A | 317 | A | 318 | A | 319 | cWW | cWW | tHH | cSH | tWH | cSs | ntHS | cWW | ntWW | ntWW | cWW | ||
| 5 | J4_9AXU_001 | 9AXU | 0.0600 | 1 | 2 | LSU rRNA | U | 112 | C | 113 | * | G | 275 | A | 276 | G | 277 | U | 278 | C | 279 | G | 280 | * | C | 301 | U | 302 | A | 303 | A | 304 | A | 305 | G | 307 | * | U | 324 | A | 325 | A | 326 | A | 327 | cWW | cWW | tHH | cSH | tWH | cSs | tHS | cWW | ntWW | ntWW | ntWW | cWW | |
| 6 | J4_7A0S_001 | 7A0S | 0.3294 | 2 | X | LSU rRNA | C | 245 | C | 246 | * | G | 383 | A | 384 | G | 385 | U | 386 | A | 387 | G | 388 | * | U | 412 | G | 413 | A | 414 | A | 415 | U | 416 | G | 419 | * | C | 434 | A | 435 | A | 436 | G | 437 | cWW | cWW | ntHH | cSH | tWH | ntHS | cWW | ntWW | ntWW | cWW | |||
| 7 | J4_4WF9_001 | 4WF9 | 0.3622 | 2 | X | LSU rRNA | C | 271 | C | 272 | * | G | 416 | A | 417 | G | 418 | U | 419 | A | 420 | C | 421 | * | G | 445 | G | 446 | A | 447 | A | 448 | U | 449 | G | 452 | * | U | 467 | A | 468 | A | 469 | G | 470 | cWW | cWW | ntHH | cSH | ntWH | cSs | tHS | cWW | ntWW | ncsS | ntWW | ncWW | |
| 8 | J4_9E6Q_001 | 9E6Q | 0.3451 | 2 | 1 | LSU rRNA | C | 249 | C | 250 | * | G | 404 | A | 405 | G | 406 | U | 407 | A | 408 | C | 409 | * | G | 430 | G | 431 | A | 432 | A | 433 | C | 434 | G | 437 | * | C | 453 | A | 454 | A | 455 | G | 456 | cWW | cWW | tHH | cSH | tWH | ncSs | tHS | cWW | ntWW | ntWW | cWW | ||
| 9 | J4_4V9F_001 | 4V9F | 0.3752 | 2 | 0 | LSU rRNA | C | 239 | C | 240 | * | G | 379 | A | 380 | G | 381 | U | 382 | A | 383 | G | 384 | * | C | 405 | G | 406 | A | 407 | A | 408 | U | 409 | C | 412 | * | G | 428 | A | 429 | A | 430 | G | 431 | cWW | cWW | tHH | cSH | tWH | cSs | tHS | cWW | ntWW | ntWW | cWW | ||
| 10 | J4_8B0X_004 | 8B0X | 0.4385 | 2 | a | LSU rRNA | C | 268 | C | 269 | * | G | 370 | A | 371 | G | 372 | U | 373 | A | 374 | G | 375 | * | U | 399 | G | 400 | A | 401 | A | 402 | U | 403 | G | 406 | * | C | 421 | A | 422 | A | 423 | G | 424 | cWW | cWW | tHH | cSH | tWH | cSs | tHS | cWW | ntWW | ntWW | ntWW | cWW | |
| 11 | J4_9DFE_001 | 9DFE | 0.3628 | 2 | 1A | LSU rRNA | C | 268 | U | 269 | * | G | 370 | A | 371 | G | 372 | U | 373 | A | 374 | C | 375 | * | G | 399 | G | 400 | A | 401 | A | 402 | U | 403 | G | 406 | * | U | 421 | A | 422 | A | 423 | G | 424 | cWW | cWW | tHH | cSH | tWH | ncSs | tHS | cWW | ntWW | ntWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UC*GAGUCG*CUAAGUG*UAAA | 2 |
| CC*GAGUCG*CCAAUCG*CAAG | 1 |
| CC*GAGUCG*CAAAGCG*UAAG | 1 |
| UC*GAGUCG*CUAAAUG*UAAA | 1 |
| CC*GAGUAG*UGAAUCCG*CAAG | 1 |
| CC*GAGUAC*GGAAUCUG*UAAG | 1 |
| CC*GAGUAC*GGAACGCG*CAAG | 1 |
| CC*GAGUAG*CGAAUAAC*GAAG | 1 |
| CC*GAGUAG*UGAAUAUG*CAAG | 1 |
| CU*GAGUAC*GGAAUCUG*UAAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| *AGUC*UAAGU*AA | 2 |
| *AGUA*GAAUCU*AA | 2 |
| *AGUC*CAAUC*AA | 1 |
| *AGUC*AAAGC*AA | 1 |
| *AGUC*UAAAU*AA | 1 |
| *AGUA*GAAUCC*AA | 1 |
| *AGUA*GAACGC*AA | 1 |
| *AGUA*GAAUAA*AA | 1 |
| *AGUA*GAAUAU*AA | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (11)
- Basepair signature
- cWW-cWW-cSS-F-tHH-cWW-cSH-tWH-F-tHS-cWW
- Heat map statistics
- Min 0.06 | Avg 0.30 | Max 0.61
Coloring options: