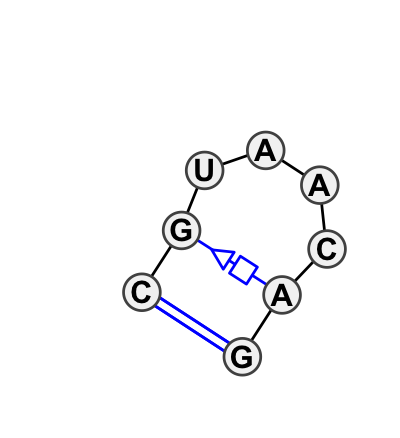
Motif HL_81943.1 Version HL_81943.1 of this group appears in releases 0.2 to 0.2
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 1-8 | 2-6 | 2-7 | 3-6 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_1QA6_002 | 1QA6 | 0.2936 | 0 | C | LSU rRNA | C | 142 | G | 143 | U | 144 | A | 145 | A | 146 | U | 147 | A | 148 | G | 149 | cWW | tSH | |||
| 2 | HL_1QA6_004 | 1QA6 | 0.1338 | 0 | D | LSU rRNA | C | 142 | G | 143 | U | 144 | A | 145 | A | 146 | U | 147 | A | 148 | G | 149 | cWW | tSH | |||
| 3 | HL_1S72_031 | 1S72 | 0.1337 | 0 | GNRA with tandem sheared (H) | 0 | LSU rRNA | C | 1196 | G | 1197 | U | 1198 | A | 1199 | A | 1200 | C | 1201 | A | 1202 | G | 1203 | cWW | tSH | ntSHa | |
| 4 | HL_1NBS_002 | 1NBS | 0.0470 | 0 | A | RNase P class B | C | 118 | U | 119 | U | 120 | U | 121 | A | 122 | G | 123 | A | 124 | G | 125 | cWW | ntSH | |||
| 5 | HL_1MMS_004 | 1MMS | 0.0000 | 0 | D | LSU rRNA | C | 1092 | G | 1093 | U | 1094 | A | 1095 | A | 1096 | C | 1097 | A | 1098 | G | 1099 | cWW | tSH | ntSHa | ||
| 6 | HL_2QBG_031 | 2QBG | 0.1285 | 0 | B | LSU rRNA | C | 1092 | G | 1093 | U | 1094 | A | 1095 | A | 1096 | U | 1097 | A | 1098 | G | 1099 | cWW | tSH | |||
| 7 | HL_1MMS_002 | 1MMS | 0.1797 | 0 | GNRA with tandem sheared | C | LSU rRNA | C | 1092 | G | 1093 | U | 1094 | A | 1095 | A | 1096 | C | 1097 | A | 1098 | G | 1099 | cWW | tSH | ntSHa | |
| 8 | HL_1MMS_001 | 1MMS | 0.3830 | 3 | C | LSU rRNA | C | 1064 | U | 1065 | U | 1066 | A | 1067 | G | 1068 | A | 1069 | A | 1073 | G | 1074 | cWW | ||||
| 9 | HL_1MMS_003 | 1MMS | 0.4608 | 3 | D | LSU rRNA | C | 1064 | U | 1065 | U | 1066 | A | 1067 | G | 1068 | A | 1069 | A | 1073 | G | 1074 | cWW | ||||
| 10 | HL_3PYO_022 | 3PYO | 0.4224 | 0 | A | LSU rRNA | G | 712 | G | 713 | U | 714 | G | 715 | A | 716 | G | 717 | A | 718 | C | 719 | cWW | ntSH | |||
| 11 | HL_1E8O_001 | 1E8O | 0.4572 | 0 | GNRA with tandem sheared | E | 7SL RNA | G | 110 | G | 111 | U | 112 | G | 113 | G | 114 | C | 115 | G | 116 | C | 117 | cWW | ntSH | ntSHa | |
| 12 | HL_2ZJR_029 | 2ZJR | 0.4572 | 0 | X | LSU rRNA | C | 1103 | G | 1104 | U | 1105 | A | 1106 | A | 1107 | U | 1108 | A | 1109 | G | 1110 | cWW | tSH | |||
| 13 | HL_1QA6_003 | 1QA6 | 0.4713 | 3 | D | LSU rRNA | C | 114 | U | 115 | U | 116 | A | 117 | G | 118 | A | 119 | A | 123 | G | 124 | cWW | ||||
| 14 | HL_2ZJR_028 | 2ZJR | 0.4596 | 3 | X | LSU rRNA | C | 1075 | U | 1076 | U | 1077 | A | 1078 | G | 1079 | A | 1080 | A | 1084 | G | 1085 | cWW | ntWH | |||
| 15 | HL_1QA6_001 | 1QA6 | 0.5195 | 3 | C | LSU rRNA | C | 114 | U | 115 | U | 116 | A | 117 | G | 118 | A | 119 | A | 123 | G | 124 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CUUAGAAGCAG | 5 |
| CGUAAUAG | 4 |
| CGUAACAG | 3 |
| CUUUAGAG | 1 |
| GGUGAGAC | 1 |
| GGUGGCGC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UUAGAAGCA | 5 |
| GUAAUA | 4 |
| GUAACA | 3 |
| UUUAGA | 1 |
| GUGAGA | 1 |
| GUGGCG | 1 |
Release history
| Release | 0.2 |
|---|---|
| Date | 2011-12-08 |
| Status | New id, 2 parents |
Parent motifs
| Parent motif | Common motif instances | Only in HL_81943.1 | Only in the parent motif |
|---|---|---|---|
| HL_70489.1 Compare | HL_1QA6_003, HL_1QA6_001, HL_1E8O_001, HL_1NBS_002, HL_2ZJR_029, HL_2ZJR_028, HL_1MMS_003, HL_1MMS_001, HL_3PYO_022 | HL_1QA6_004, HL_1QA6_002, HL_1S72_031, HL_2QBG_031, HL_1MMS_002, HL_1MMS_004 | HL_2QBG_023, HL_1S72_030, HL_3PYO_057, HL_2ZJR_025, HL_3O58_050, HL_2QBG_051 |
| HL_79793.1 Compare | HL_1QA6_004, HL_1QA6_002, HL_1S72_031, HL_2QBG_031, HL_1MMS_002, HL_1MMS_004 | HL_1QA6_003, HL_1QA6_001, HL_1E8O_001, HL_1NBS_002, HL_2ZJR_029, HL_2ZJR_028, HL_1MMS_003, HL_1MMS_001, HL_3PYO_022 | HL_1OOA_002 |
Children motifs
| Child motif | Common motif instances | Only in HL_81943.1 | Only in the child motif |
|---|---|---|---|
| HL_18781.1 Compare | HL_1QA6_002, HL_1S72_031, HL_2QBG_031, HL_1E8O_001, HL_1NBS_002, HL_2ZJR_029 | HL_1QA6_004, HL_1QA6_003, HL_1QA6_001, HL_2ZJR_028, HL_1MMS_003, HL_1MMS_002, HL_1MMS_001, HL_3PYO_022, HL_1MMS_004 | HL_2QBG_023, HL_1U9S_003, HL_4A1B_032, HL_2ZJR_055, HL_3U5H_028, HL_3NPB_003, HL_2XZM_008, HL_3V2F_055, HL_3U5F_008, HL_1FJG_006, HL_3V2F_022, HL_4A1B_012, HL_2IL9_006, HL_3Q1Q_005, HL_2AW7_006 |
| HL_72543.1 Compare | HL_1QA6_001, HL_2ZJR_028 | HL_1QA6_004, HL_1QA6_002, HL_1QA6_003, HL_1S72_031, HL_2QBG_031, HL_1E8O_001, HL_1NBS_002, HL_2ZJR_029, HL_1MMS_003, HL_1MMS_002, HL_1MMS_001, HL_3PYO_022, HL_1MMS_004 | HL_1S72_030, HL_2QBG_030, HL_3U5H_027 |
- Annotations
-
- (12)
- GNRA with tandem sheared (2)
- GNRA with tandem sheared (H) (1)
- Basepair signature
- Heat map statistics
- Min 0.00 | Avg 0.36 | Max 0.56
Coloring options: