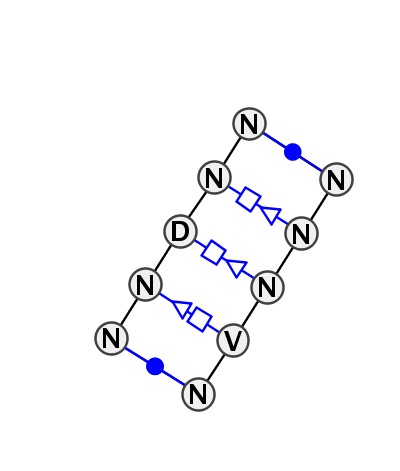
Motif IL_35765.1 Version IL_35765.1 of this group appears in releases 2.0 to 2.0
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | break | 6 | 7 | 8 | 9 | 10 | 1-10 | 2-3 | 2-9 | 3-8 | 4-7 | 5-6 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_4V9F_020 | 4V9F | 0.5997 | 0 | AAA cross-strand stack | 0 | LSU rRNA | C | 663 | U | 664 | A | 665 | A | 666 | C | 667 | * | G | 679 | G | 680 | G | 681 | A | 682 | G | 683 | cWW | ntWH | tHS | cWW | ||
| 2 | IL_5J7L_258 | 5J7L | 0.6126 | 0 | AAA cross-strand stack | DA | LSU rRNA | U | 606 | U | 607 | A | 608 | A | 609 | C | 610 | * | G | 618 | G | 619 | G | 620 | A | 621 | G | 622 | cWW | ntWH | tHS | cWW | ||
| 3 | IL_5FDU_016 | 5FDU | 0.5743 | 0 | AAA cross-strand stack (H) | 1A | LSU rRNA | U | 629 | U | 630 | A | 631 | A | 632 | G | 633 | * | C | 643 | G | 644 | G | 645 | A | 646 | G | 647 | cWW | ntWH | tHS | cWW | ||
| 4 | IL_4YAZ_006 | 4YAZ | 0.5070 | 0 | Triple sheared | R | Cyclic di-GMP-I riboswitch | U | 16 | A | 17 | A | 18 | A | 19 | C | 20 | * | G | 35 | G | 36 | G | 37 | C | 38 | G | 39 | cWW | ntSH | tHS | cWW | ||
| 5 | IL_3UCZ_001 | 3UCZ | 0.4781 | 0 | R | Cyclic di-GMP-I riboswitch | C | 22 | A | 23 | A | 24 | A | 25 | C | 26 | * | G | 41 | G | 42 | A | 43 | C | 44 | G | 45 | cWW | ntSH | tHS | cWW | |||
| 6 | IL_4V91_034 | 4V91 | 0.4643 | 0 | 1 | LSU rRNA | U | 834 | G | 835 | A | 836 | A | 837 | G | 838 | * | U | 855 | G | 856 | G | 857 | A | 858 | G | 859 | cWW | ntSH | ntHS | tHS | cWW | ||
| 7 | IL_4V88_273 | 4V88 | 0.4512 | 0 | Triple sheared (H) | A5 | LSU rRNA | U | 834 | G | 835 | A | 836 | A | 837 | G | 838 | * | U | 855 | G | 856 | G | 857 | A | 858 | G | 859 | cWW | tSH | tHS | tHS | cWW | |
| 8 | IL_3J79_032 | 3J79 | 0.4395 | 0 | A | LSU rRNA | U | 953 | G | 954 | A | 955 | A | 956 | G | 957 | * | U | 974 | G | 975 | G | 976 | A | 977 | G | 978 | cWW | ntSH | tHS | tHS | cWW | ||
| 9 | IL_5J7L_263 | 5J7L | 0.4358 | 0 | Triple sheared | DA | LSU rRNA | U | 703 | G | 704 | A | 705 | A | 706 | G | 707 | * | U | 724 | G | 725 | G | 726 | A | 727 | G | 728 | cWW | tSH | tHS | tHS | cWW | |
| 10 | IL_5FDU_021 | 5FDU | 0.4208 | 0 | Triple sheared (H) | 1A | LSU rRNA | U | 750 | G | 751 | A | 752 | A | 753 | G | 754 | * | U | 771 | G | 772 | G | 773 | A | 774 | G | 775 | cWW | tSH | tHS | tHS | cWW | |
| 11 | IL_4V9F_027 | 4V9F | 0.4170 | 0 | Triple sheared | 0 | LSU rRNA | U | 794 | G | 795 | A | 796 | A | 797 | G | 798 | * | U | 815 | G | 816 | G | 817 | A | 818 | A | 819 | cWW | ntSH | tHS | tHS | cWW | |
| 12 | IL_5AJ0_063 | 5AJ0 | 0.4365 | 0 | Triple sheared (H) | A2 | LSU rRNA | C | 1533 | G | 1534 | A | 1535 | A | 1536 | G | 1537 | * | U | 1554 | G | 1555 | G | 1556 | A | 1557 | G | 1558 | cWW | ntSH | tHS | tHS | cWW | |
| 13 | IL_3J9W_086 | 3J9W | 0.3617 | 0 | BA | LSU rRNA | U | 750 | G | 751 | A | 752 | A | 753 | G | 754 | * | U | 771 | G | 772 | G | 773 | A | 774 | G | 775 | cWW | tSH | tHS | tHS | cWW | ||
| 14 | IL_5J7L_310 | 5J7L | 0.3619 | 0 | Triple sheared | DA | LSU rRNA | U | 1720 | G | 1721 | A | 1722 | G | 1723 | G | 1724 | * | U | 1736 | G | 1737 | G | 1738 | A | 1739 | G | 1740 | cWW | tSH | tHS | ntHS | cWW | |
| 15 | IL_3IWN_002 | 3IWN | 0.4110 | 0 | Triple sheared | A | Cyclic di-GMP-I riboswitch | C | 12 | A | 13 | A | 14 | A | 15 | C | 16 | * | G | 31 | G | 32 | A | 33 | C | 34 | G | 35 | cWW | ntSH | tHS | cWW | ||
| 16 | IL_3J79_064 | 3J79 | 0.4163 | 0 | Triple sheared (H) | A | LSU rRNA | U | 1800 | G | 1801 | A | 1802 | A | 1803 | C | 1804 | * | G | 2028 | G | 2029 | G | 2030 | A | 2031 | A | 2032 | cWW | tSH | tHS | tHS | cWW | |
| 17 | IL_4V91_072 | 4V91 | 0.4236 | 0 | Triple sheared (H) | 1 | LSU rRNA | U | 1645 | G | 1646 | A | 1647 | A | 1648 | U | 1649 | * | A | 1806 | G | 1807 | G | 1808 | A | 1809 | A | 1810 | cWW | tSH | tHS | tHS | cWW | |
| 18 | IL_4V88_316 | 4V88 | 0.3614 | 0 | Triple sheared (H) | A5 | LSU rRNA | U | 1645 | G | 1646 | A | 1647 | A | 1648 | U | 1649 | * | A | 1806 | G | 1807 | G | 1808 | A | 1809 | A | 1810 | cWW | tSH | tHS | tHS | cWW | |
| 19 | IL_3J9W_142 | 3J9W | 0.3008 | 0 | Triple sheared (H) | BA | LSU rRNA | C | 1911 | G | 1912 | A | 1913 | A | 1914 | U | 1915 | * | A | 1885 | G | 1886 | G | 1887 | A | 1888 | G | 1889 | cWW | tSH | tHS | tHS | cWW | |
| 20 | IL_5AJ0_106 | 5AJ0 | 0.3684 | 0 | Triple sheared (H) | A2 | LSU rRNA | C | 2568 | G | 2569 | A | 2570 | U | 2571 | C | 2572 | * | G | 2731 | G | 2732 | G | 2733 | A | 2734 | G | 2735 | cWW | tSH | tHS | ntHS | cWW | |
| 21 | IL_3J7P_085 | 3J7P | 0.2999 | 0 | Triple sheared (H) | 5 | LSU rRNA | C | 2589 | G | 2590 | A | 2591 | U | 2592 | C | 2593 | * | G | 2752 | G | 2753 | G | 2754 | A | 2755 | G | 2756 | cWW | tSH | ntHS | ntHS | cWW | |
| 22 | IL_2A64_006 | 2A64 | 0.3372 | 0 | Triple sheared | A | RNase P class B | C | 378 | G | 379 | A | 380 | A | 381 | G | 382 | * | C | 347 | G | 348 | G | 349 | A | 350 | G | 351 | cWW | tHS | tHS | cWW | ||
| 23 | IL_4IOA_070 | 4IOA | 0.3275 | 0 | Triple sheared | X | LSU rRNA | C | 1865 | G | 1866 | A | 1867 | A | 1868 | A | 1869 | * | U | 1848 | G | 1849 | G | 1850 | A | 1851 | G | 1852 | cWW | ntSH | tHS | tHS | cWW | |
| 24 | IL_5FDU_076 | 5FDU | 0.3193 | 0 | Triple sheared (H) | 1A | LSU rRNA | C | 1904 | G | 1905 | A | 1906 | A | 1907 | C | 1908 | * | G | 1887 | G | 1888 | G | 1889 | A | 1890 | G | 1891 | cWW | tSH | tHS | tHS | cWW | |
| 25 | IL_5A2Q_089 | 5A2Q | 0.2944 | 0 | Triple sheared (H) | 2 | SSU rRNA | A | 1788 | G | 1789 | A | 1790 | A | 1791 | G | 1792 | * | C | 1742 | G | 1743 | G | 1744 | A | 1745 | U | 1746 | cWW | ntSH | ntHS | tHS | cWW | |
| 26 | IL_4BTS_349 | 4BTS | 0.3200 | 0 | Triple sheared (H) | DA | SSU rRNA | G | 1672 | A | 1673 | A | 1674 | A | 1675 | A | 1676 | * | U | 1649 | G | 1650 | G | 1651 | A | 1652 | C | 1653 | cWW | tSH | tHS | tHS | cWW | |
| 27 | IL_2A64_005 | 2A64 | 0.3133 | 0 | Triple sheared | A | RNase P class B | C | 284 | G | 285 | A | 286 | A | 287 | G | 288 | * | C | 297 | G | 298 | G | 299 | A | 300 | G | 301 | cWW | tSH | ntHS | tHS | cWW | |
| 28 | IL_3DHS_004 | 3DHS | 0.3860 | 0 | Triple sheared | A | RNase P class B | C | 284 | G | 285 | A | 286 | A | 287 | G | 288 | * | C | 297 | G | 298 | G | 299 | A | 300 | G | 301 | cWW | ncSH | tHS | cWW | ||
| 29 | IL_4V8P_427 | 4V8P | 0.4261 | 0 | F1 | LSU rRNA | U | 3217 | U | 3218 | A | 3219 | A | 3220 | C | 3221 | * | G | 3189 | A | 3190 | G | 3191 | C | 3192 | G | 3193 | cWW | ntSHa | ntHS | tHS | cWW | ||
| 30 | IL_3JAM_085 | 3JAM | 0.4381 | 0 | 2 | SSU rRNA | A | 1717 | G | 1718 | A | 1719 | A | 1720 | U | 1721 | * | A | 1676 | G | 1677 | G | 1678 | A | 1679 | U | 1680 | cWW | ntsS | tHS | tHS | cWW | ||
| 31 | IL_4V88_474 | 4V88 | 0.4394 | 0 | Triple sheared | A6 | SSU rRNA | A | 1719 | G | 1720 | A | 1721 | A | 1722 | U | 1723 | * | A | 1678 | G | 1679 | G | 1680 | A | 1681 | U | 1682 | cWW | tHS | tHS | cWW | ||
| 32 | IL_3J7A_080 | 3J7A | 0.4331 | 0 | Triple sheared (H) | A | SSU rRNA | G | 2011 | G | 2012 | A | 2013 | A | 2014 | A | 2015 | * | U | 1975 | G | 1976 | G | 1977 | A | 1978 | C | 1979 | cWW | tHS | tHS | cWW | ||
| 33 | IL_1U6B_002 | 1U6B | 0.2713 | 0 | Double sheared with non-canonical cWW | B | 197-MER | G | 56 | A | 57 | A | 58 | A | 59 | C | 60 | * | G | 84 | C | 85 | A | 86 | A | 87 | C | 88 | cWW | tSH | tHS | ncWW | cWW | |
| 34 | IL_3J7P_233 | 3J7P | 0.2530 | 0 | S2 | SSU rRNA | U | 1733 | G | 1734 | A | 1735 | G | 1736 | G | 1737 | * | U | 1797 | C | 1798 | G | 1799 | A | 1800 | A | 1801 | cWW | tSH | tHS | ncWW | cWW | ||
| 35 | IL_4V9F_085 | 4V9F | 0.2911 | 0 | Double sheared with non-canonical cWW | 0 | LSU rRNA | C | 2515 | G | 2516 | A | 2517 | C | 2518 | C | 2519 | * | G | 2501 | C | 2502 | A | 2503 | A | 2504 | G | 2505 | cWW | tSH | ntHS | cBW | cWW | |
| 36 | IL_4V4Q_235 | 4V4Q | 0.3594 | 0 | Double sheared with non-canonical cWW (H) | CA | SSU rRNA | C | 1259 | G | 1260 | A | 1261 | C | 1262 | C | 1263 | * | G | 1272 | C | 1273 | A | 1274 | A | 1275 | G | 1276 | cWW | tSH | tHS | ncBW | cWW | |
| 37 | IL_3J9W_050 | 3J9W | 0.2551 | 0 | Double sheared with non-canonical cWW (H) | AA | SSU rRNA | C | 1268 | G | 1269 | A | 1270 | A | 1271 | A | 1272 | * | U | 1281 | U | 1282 | A | 1283 | A | 1284 | G | 1285 | cWW | tSH | tHS | cWW | ||
| 38 | IL_5AJ3_006 | 5AJ3 | 0.2161 | 0 | A | SSU rRNA | C | 97 | A | 98 | A | 99 | A | 100 | A | 101 | * | U | 81 | C | 82 | C | 83 | A | 84 | G | 85 | cWW | tSH | tHS | cWW | |||
| 39 | IL_5J7L_274 | 5J7L | 0.1808 | 0 | Double sheared with non-canonical cWW | DA | LSU rRNA | C | 898 | A | 899 | A | 900 | C | 901 | C | 902 | * | G | 875 | C | 876 | A | 877 | A | 878 | G | 879 | cWW | tSH | ntHS | cBW | cWW | |
| 40 | IL_4WFL_001 | 4WFL | 0.0000 | 0 | Double sheared with non-canonical cWW | A | LSRP RNA | C | 91 | G | 92 | A | 93 | C | 94 | A | 95 | * | U | 72 | C | 73 | G | 74 | A | 75 | G | 76 | cWW | tSH | tHS | cBW | cWW | |
| 41 | IL_3J7Y_046 | 3J7Y | 0.2014 | 0 | A | LSU rRNA | C | 2872 | A | 2873 | A | 2874 | A | 2875 | G | 2876 | * | C | 2856 | U | 2857 | A | 2858 | A | 2859 | G | 2860 | cWW | tSH | tHS | cWW | |||
| 42 | IL_3J9W_164 | 3J9W | 0.2140 | 0 | Double sheared with non-canonical cWW (H) | BA | LSU rRNA | C | 2509 | G | 2510 | A | 2511 | C | 2512 | G | 2513 | * | C | 2495 | C | 2496 | A | 2497 | A | 2498 | G | 2499 | cWW | tSH | tHS | ncBW | cWW | |
| 43 | IL_5J7L_338 | 5J7L | 0.1643 | 0 | Double sheared with non-canonical cWW | DA | LSU rRNA | C | 2480 | G | 2481 | A | 2482 | C | 2483 | G | 2484 | * | C | 2466 | C | 2467 | A | 2468 | A | 2469 | G | 2470 | cWW | tSH | ntHS | cBW | cWW | |
| 44 | IL_4V8P_412 | 4V8P | 0.2059 | 0 | F1 | LSU rRNA | C | 2837 | G | 2838 | A | 2839 | C | 2840 | G | 2841 | * | C | 2823 | C | 2824 | A | 2825 | A | 2826 | G | 2827 | cWW | tSH | tHS | ncBW | cWW | ||
| 45 | IL_5AJ0_150 | 5AJ0 | 0.2031 | 0 | A2 | LSU rRNA | C | 4388 | G | 4389 | A | 4390 | C | 4391 | G | 4392 | * | C | 4374 | C | 4375 | A | 4376 | A | 4377 | G | 4378 | cWW | tSH | ncBW | cWW | |||
| 46 | IL_4V19_048 | 4V19 | 0.2640 | 0 | Double sheared with non-canonical cWW (H) | A | LSU rRNA | C | 1301 | G | 1302 | A | 1303 | C | 1304 | A | 1305 | * | U | 1287 | C | 1288 | U | 1289 | A | 1290 | G | 1291 | cWW | ntSH | ncBW | cWW | ||
| 47 | IL_4IOA_089 | 4IOA | 0.2444 | 0 | Triple sheared | X | LSU rRNA | C | 2459 | G | 2460 | G | 2461 | C | 2462 | G | 2463 | * | C | 2445 | C | 2446 | G | 2447 | A | 2448 | G | 2449 | cWW | tSH | ntHS | ncBW | cWW | |
| 48 | IL_5FDU_097 | 5FDU | 0.2612 | 0 | Triple sheared (H) | 1A | LSU rRNA | C | 2492 | G | 2493 | G | 2494 | C | 2495 | G | 2496 | * | C | 2478 | C | 2479 | G | 2480 | A | 2481 | G | 2482 | cWW | tSH | cBW | cWW | ||
| 49 | IL_3J79_103 | 3J79 | 0.3328 | 0 | A | LSU rRNA | C | 3208 | G | 3209 | A | 3210 | C | 3211 | G | 3212 | * | C | 3194 | C | 3195 | A | 3196 | A | 3197 | G | 3198 | cWW | tSH | ntHS | cWW | |||
| 50 | IL_4V88_353 | 4V88 | 0.3670 | 0 | Triple sheared (H) | A5 | LSU rRNA | C | 2849 | G | 2850 | A | 2851 | C | 2852 | A | 2853 | * | U | 2835 | C | 2836 | A | 2837 | A | 2838 | G | 2839 | cWW | tSH | ntHS | cWW | ||
| 51 | IL_4V91_115 | 4V91 | 0.3673 | 0 | Triple sheared (H) | 1 | LSU rRNA | C | 2849 | G | 2850 | A | 2851 | C | 2852 | A | 2853 | * | U | 2835 | C | 2836 | A | 2837 | A | 2838 | G | 2839 | cWW | tSH | ntHS | cWW | ||
| 52 | IL_5AN9_039 | 5AN9 | 0.3967 | 0 | N | LSU rRNA | C | 3182 | G | 3183 | A | 3184 | C | 3185 | G | 3186 | * | C | 3168 | C | 3169 | A | 3170 | A | 3171 | G | 3172 | cWW | tSH | ntHS | cWW | |||
| 53 | IL_3J7P_122 | 3J7P | 0.3720 | 0 | Triple sheared (H) | 5 | LSU rRNA | C | 4426 | G | 4427 | A | 4428 | C | 4429 | G | 4430 | * | C | 4412 | C | 4413 | A | 4414 | A | 4415 | G | 4416 | ncWW | tSH | tHS | cWW | ||
| 54 | IL_5AJ0_056 | 5AJ0 | 0.3475 | 0 | A2 | LSU rRNA | C | 1432 | C | 1433 | G | 1434 | A | 1435 | G | 1436 | * | C | 1413 | C | 1414 | G | 1415 | A | 1416 | G | 1417 | ncWW | tSH | tHS | cWW | cWW | ||
| 55 | IL_3J7P_168 | 3J7P | 0.3983 | 0 | S2 | SSU rRNA | G | 207 | G | 208 | A | 209 | U | 210 | G | 211 | * | C | 188 | U | 189 | G | 190 | A | 191 | C | 192 | cWW | tSH | ntHS | ncwW | cWW | ||
| 56 | IL_3J7P_223 | 3J7P | 0.5860 | 0 | S2 | SSU rRNA | C | 1559 | U | 1560 | A | 1561 | C | 1562 | G | 1563 | * | C | 1572 | G | 1573 | C | 1574 | G | 1575 | G | 1576 | cWW | ncWW | cWW | ||||
| 57 | IL_3J7A_061 | 3J7A | 0.7444 | 1 | A | SSU rRNA | C | 1440 | C | 1441 | U | 1442 | C | 1444 | U | 1445 | * | A | 1628 | G | 1629 | A | 1630 | G | 1631 | G | 1632 | cWW | ncWW | cWW | ||||
| 58 | IL_3J7A_047 | 3J7A | 0.7122 | 0 | tSH-tWH-tHS (H) | A | SSU rRNA | U | 1059 | G | 1060 | A | 1061 | A | 1062 | G | 1063 | * | C | 1079 | G | 1080 | U | 1081 | A | 1082 | A | 1083 | cWW | tSH | tHS | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CGACG*CCAAG | 7 |
| UGAAG*UGGAG | 6 |
| CGAAG*CGGAG | 3 |
| CAAAC*GGACG | 2 |
| UGAAU*AGGAA | 2 |
| CGAUC*GGGAG | 2 |
| AGAAU*AGGAU | 2 |
| CGACC*GCAAG | 2 |
| CGGCG*CCGAG | 2 |
| CGACA*UCAAG | 2 |
| CUAAC*GGGAG | 1 |
| UUAAC*GGGAG | 1 |
| UUAAG*CGGAG | 1 |
| UAAAC*GGGCG | 1 |
| UGAAG*UGGAA | 1 |
| CGAAG*UGGAG | 1 |
| UGAGG*UGGAG | 1 |
| UGAAC*GGGAA | 1 |
| CGAAU*AGGAG | 1 |
| CGAAA*UGGAG | 1 |
| CGAAC*GGGAG | 1 |
| AGAAG*CGGAU | 1 |
| GAAAA*UGGAC | 1 |
| UUAAC*GAGCG | 1 |
| GGAAA*UGGAC | 1 |
| GAAAC*GCAAC | 1 |
| UGAGG*UCGAA | 1 |
| CGAAA*UUAAG | 1 |
| CAAAA*UCCAG | 1 |
| CAACC*GCAAG | 1 |
| CGACA*UCGAG | 1 |
| CAAAG*CUAAG | 1 |
| CGACA*UCUAG | 1 |
| CCGAG*CCGAG | 1 |
| GGAUG*CUGAC | 1 |
| CUACG*CGCGG | 1 |
| CCUGCU*AGAGG | 1 |
| UGAAG*CGUAA | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAA*GGA | 21 |
| GAC*CAA | 11 |
| UAA*GGA | 3 |
| AAA*GAC | 2 |
| GAU*GGA | 2 |
| GGC*CGA | 2 |
| AAA*GGC | 1 |
| GAG*GGA | 1 |
| AAA*GGA | 1 |
| UAA*AGC | 1 |
| AAA*CAA | 1 |
| GAG*CGA | 1 |
| GAA*UAA | 1 |
| AAA*CCA | 1 |
| AAC*CAA | 1 |
| GAC*CGA | 1 |
| AAA*UAA | 1 |
| GAC*CUA | 1 |
| CGA*CGA | 1 |
| GAU*UGA | 1 |
| UAC*GCG | 1 |
| CUGC*GAG | 1 |
| GAA*GUA | 1 |
Release history
| Release | 2.0 |
|---|---|
| Date | 2017-04-24 |
| Status | New id, 2 parents |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (17)
- Triple sheared (H) (17)
- Triple sheared (11)
- Double sheared with non-canonical cWW (5)
- Double sheared with non-canonical cWW (H) (4)
- AAA cross-strand stack (2)
- tSH-tWH-tHS (H) (1)
- AAA cross-strand stack (H) (1)
- Basepair signature
- cWW-tSH-tHS-tHS-cWW
- Heat map statistics
- Min 0.06 | Avg 0.47 | Max 1.01
Coloring options: